Juvenile hormones direct primordial germ cell migration to the embryonic gonad
- PMID: 38215744
- PMCID: PMC10872347
- DOI: 10.1016/j.cub.2023.12.033
Juvenile hormones direct primordial germ cell migration to the embryonic gonad
Abstract
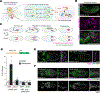
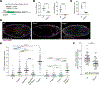
Germ cells are essential to sexual reproduction. Across the animal kingdom, extracellular signaling isoprenoids, such as retinoic acids (RAs) in vertebrates and juvenile hormones (JHs) in invertebrates, facilitate multiple processes in reproduction. Here we investigated the role of these potent signaling molecules in embryonic germ cell development, using JHs in Drosophila melanogaster as a model system. In contrast to their established endocrine roles during larval and adult germline development, we found that JH signaling acts locally during embryonic development. Using an in vivo biosensor, we observed active JH signaling first within and near primordial germ cells (PGCs) as they migrate to the developing gonad. Through in vivo and in vitro assays, we determined that JHs are both necessary and sufficient for PGC migration. Analysis into the mechanisms of this newly uncovered paracrine JH function revealed that PGC migration was compromised when JHs were decreased or increased, suggesting that specific titers or spatiotemporal JH dynamics are required for robust PGC colonization of the gonad. Compromised PGC migration can impair fertility and cause germ cell tumors in many species, including humans. In mammals, retinoids have many roles in development and reproduction. We found that like JHs in Drosophila, RA was sufficient to impact mouse PGC migration in vitro. Together, our study reveals a previously unanticipated role of isoprenoids as local effectors of pre-gonadal PGC development and suggests a broadly shared mechanism in PGC migration.
Keywords: Hmgcr; cell movement; embryonic development; gametogenesis; germ cells; gonad; juvenile hormones; ovary; retinoids; testis.
Copyright © 2023 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures




Comment in
-
Germ cell migration: Unexpected role of juvenile hormone before juvenile stages.Curr Biol. 2024 Feb 5;34(3):R84-R86. doi: 10.1016/j.cub.2023.12.026. Curr Biol. 2024. PMID: 38320477
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

