Hagfish genome elucidates vertebrate whole-genome duplication events and their evolutionary consequences
- PMID: 38216617
- PMCID: PMC10927551
- DOI: 10.1038/s41559-023-02299-z
Hagfish genome elucidates vertebrate whole-genome duplication events and their evolutionary consequences
Abstract
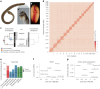
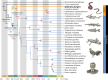
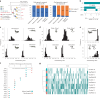
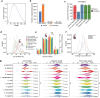
Polyploidy or whole-genome duplication (WGD) is a major event that drastically reshapes genome architecture and is often assumed to be causally associated with organismal innovations and radiations. The 2R hypothesis suggests that two WGD events (1R and 2R) occurred during early vertebrate evolution. However, the timing of the 2R event relative to the divergence of gnathostomes (jawed vertebrates) and cyclostomes (jawless hagfishes and lampreys) is unresolved and whether these WGD events underlie vertebrate phenotypic diversification remains elusive. Here we present the genome of the inshore hagfish, Eptatretus burgeri. Through comparative analysis with lamprey and gnathostome genomes, we reconstruct the early events in cyclostome genome evolution, leveraging insights into the ancestral vertebrate genome. Genome-wide synteny and phylogenetic analyses support a scenario in which 1R occurred in the vertebrate stem-lineage during the early Cambrian, and 2R occurred in the gnathostome stem-lineage, maximally in the late Cambrian-earliest Ordovician, after its divergence from cyclostomes. We find that the genome of stem-cyclostomes experienced an additional independent genome triplication. Functional genomic and morphospace analyses demonstrate that WGD events generally contribute to developmental evolution with similar changes in the regulatory genome of both vertebrate groups. However, appreciable morphological diversification occurred only in the gnathostome but not in the cyclostome lineage, calling into question the general expectation that WGDs lead to leaps of bodyplan complexity.
© 2024. The Author(s).
Conflict of interest statement
M.D.C. is a cofounder and shareholder of NovAb, Inc., which produces lamprey antibodies for biomedical purposes. J.P.R. is a consultant for NovAb. The other authors declare no competing interests.
Figures










References
-
- Van de Peer Y, Maere S, Meyer A. The evolutionary significance of ancient genome duplications. Nat. Rev. Genet. 2009;10:725–732. - PubMed
-
- Ohno, S. Evolution by Gene Duplication (Springer, 1970).
-
- Donoghue P, Purnell M. Genome duplication, extinction and vertebrate evolution. Trends Ecol. Evol. 2005;20:312–319. - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

