Identification of sulfur metabolism-related gene signature in osteoarthritis and TM9SF2's sustenance effect on M2 macrophages' phagocytic activity
- PMID: 38218914
- PMCID: PMC10787471
- DOI: 10.1186/s13018-023-04384-2
Identification of sulfur metabolism-related gene signature in osteoarthritis and TM9SF2's sustenance effect on M2 macrophages' phagocytic activity
Abstract
Background: Osteoarthritis (OA) is a chronic and low-grade inflammatory disease associated with metabolism disorder and multiple cell death types in the synovial tissues. Sulfur metabolism has not been studied in OA.
Methods: First, we calculated the single sample gene set enrichment analysis score of sulfur metabolism-associated annotations (i.e., cysteine metabolism process, regulation of sulfur metabolism process, and disulfidptosis) between healthy and synovial samples from patients with OA. Sulfur metabolism-related differentially expressed genes (DEGs) were analyzed in OA. Least absolute shrinkage and selection operator COX regression were used to identify the sulfur metabolism-associated gene signature for diagnosing OA. Correlation and immune cell deconvolution analyses were used to explore the correlated functions and cell specificity of the signature gene, TM9SF2. TM9SF2's effect on the phagocytosis of macrophages M2 was analyzed by coculturing macrophages with IgG-coated beads or apoptotic Jurkat cells.
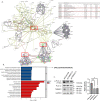
Results: A diagnostic six gene signature (i.e., MTHFD1, PDK4, TM9SF2, POU4F1, HOXA2, NCKAP1) was identified based on the ten DEGs, validated using GSE12021 and GSE1919 datasets. TM9SF2 was upregulated in the synovial tissues of OA at both mRNA and protein levels. The relationship between TM9SF2 and several functional annotations, such as antigen processing and presentation, lysosome, phagosome, Fcγ-mediated phagocytosis, and tyrosine metabolism, was identified. TM9SF2 and macrophages M2 were significantly correlated. After silencing TM9SF2 in THP-1-derived macrophages M2, a significantly reduced phagocytosis and attenuated activation of PLC-γ1 were observed.
Conclusion: A sulfur metabolism-associated six-gene signature for OA diagnosis was constructed and upregulation of the phagocytosis-associated gene, TM9SF2, was identified. The findings are expected to deepen our understanding of the molecular mechanism underlying OA development and be used as potential therapeutic targets.
Keywords: Diagnosis; Osteoarthritis; Phagocytosis; Sulfur metabolism; TM9SF2.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






Similar articles
-
Comprehensive multiomics analysis identifies PYCARD as a key pyroptosis-related gene in osteoarthritis synovial macrophages.Front Immunol. 2025 Mar 24;16:1558139. doi: 10.3389/fimmu.2025.1558139. eCollection 2025. Front Immunol. 2025. PMID: 40196125 Free PMC article.
-
Bioinformatics analysis to identify key genes and pathways influencing synovial inflammation in osteoarthritis.Mol Med Rep. 2018 Dec;18(6):5594-5602. doi: 10.3892/mmr.2018.9575. Epub 2018 Oct 23. Mol Med Rep. 2018. PMID: 30365099 Free PMC article.
-
Comprehensive bulk and single-cell transcriptome profiling give useful insights into the characteristics of osteoarthritis associated synovial macrophages.Front Immunol. 2023 Jan 5;13:1078414. doi: 10.3389/fimmu.2022.1078414. eCollection 2022. Front Immunol. 2023. PMID: 36685529 Free PMC article.
-
Effects of synovial macrophages in osteoarthritis.Front Immunol. 2023 Jul 10;14:1164137. doi: 10.3389/fimmu.2023.1164137. eCollection 2023. Front Immunol. 2023. PMID: 37492583 Free PMC article. Review.
-
Macrophages regulate the progression of osteoarthritis.Osteoarthritis Cartilage. 2020 May;28(5):555-561. doi: 10.1016/j.joca.2020.01.007. Epub 2020 Jan 23. Osteoarthritis Cartilage. 2020. PMID: 31982565 Review.
Cited by
-
A comparative metabolomic analysis reveals the metabolic variations among cartilage of Kashin-Beck disease and osteoarthritis.Bone Joint Res. 2024 Jul 17;13(7):362-371. doi: 10.1302/2046-3758.137.BJR-2023-0403.R1. Bone Joint Res. 2024. PMID: 39013544 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

