Small Molecule Screen Identifies Non-catalytic USP3 Chemical Handle
- PMID: 38222562
- PMCID: PMC10785082
- DOI: 10.1021/acsomega.3c07070
Small Molecule Screen Identifies Non-catalytic USP3 Chemical Handle
Abstract
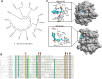
Zinc-finger ubiquitin-binding domains (ZnF-UBDs) are noncatalytic domains mostly found in deubiquitylases (DUBs) such as USP3. They represent an underexplored opportunity for the development of deubiquitylase-targeting chimeras (DUBTACs) to pharmacologically induce the deubiquitylation of target proteins. We previously showed that ZnF-UBDs are ligandable domains. Here, a focused small molecule library screen against a panel of 11 ZnF-UBDs led to the identification of compound 59, a ligand engaging the ZnF-UBD of USP3 with a KD of 14 μM. The compound binds the expected C-terminal ubiquitin binding pocket of USP3 as shown by hydrogen-deuterium exchange mass spectrometry experiments and does not inhibit the cleavage of K48-linked diubiquitin by USP3. As such, this molecule is a chemical starting point toward chemical tools that could be used to interrogate the function of the USP3 Znf-UBD and the consequences of recruiting USP3 to ubiquitylated proteins.
© 2023 The Authors. Published by American Chemical Society.
Conflict of interest statement
The authors declare no competing financial interest.
Figures



Similar articles
-
Structure-Activity Relationship of USP5 Inhibitors.J Med Chem. 2021 Oct 28;64(20):15017-15036. doi: 10.1021/acs.jmedchem.1c00889. Epub 2021 Oct 14. J Med Chem. 2021. PMID: 34648286
-
Identification and Structure-Activity Relationship of HDAC6 Zinc-Finger Ubiquitin Binding Domain Inhibitors.J Med Chem. 2018 May 24;61(10):4517-4527. doi: 10.1021/acs.jmedchem.8b00258. Epub 2018 May 9. J Med Chem. 2018. PMID: 29741882
-
USP3 inhibits type I interferon signaling by deubiquitinating RIG-I-like receptors.Cell Res. 2014 Apr;24(4):400-16. doi: 10.1038/cr.2013.170. Epub 2013 Dec 24. Cell Res. 2014. PMID: 24366338 Free PMC article.
-
The 'dark matter' of ubiquitin-mediated processes: opportunities and challenges in the identification of ubiquitin-binding domains.Biochem Soc Trans. 2019 Dec 20;47(6):1949-1962. doi: 10.1042/BST20190869. Biochem Soc Trans. 2019. PMID: 31829417 Review.
-
Ubiquitin specific peptidase 3: an emerging deubiquitinase that regulates physiology and diseases.Cell Death Discov. 2024 May 21;10(1):243. doi: 10.1038/s41420-024-02010-6. Cell Death Discov. 2024. PMID: 38773075 Free PMC article. Review.
Cited by
-
A Target Class Ligandability Evaluation of WD40 Repeat-Containing Proteins.J Med Chem. 2025 Jan 23;68(2):1092-1112. doi: 10.1021/acs.jmedchem.4c02010. Epub 2024 Nov 4. J Med Chem. 2025. PMID: 39495097 Free PMC article.
-
Quantitative Hydrogen-Deuterium Exchange Mass Spectrometry for Simultaneous Structural Characterization and Affinity Indexing of Single Target Drug Candidate Libraries.Anal Chem. 2024 Aug 13;96(32):13015-13024. doi: 10.1021/acs.analchem.4c01001. Epub 2024 Jul 29. Anal Chem. 2024. PMID: 39074309 Free PMC article.
References
-
- Henning N. J.; Boike L.; Spradlin J. N.; Ward C. C.; Liu G.; Zhang E.; Belcher B. P.; Brittain S. M.; Hesse M. J.; Dovala D.; McGregor L. M.; Valdez Misiolek R.; Plasschaert L. W.; Rowlands D. J.; Wang F.; Frank A. O.; Fuller D.; Estes A. R.; Randal K. L.; Panidapu A.; McKenna J. M.; Tallarico J. A.; Schirle M.; Nomura D. K. Deubiquitinase-Targeting Chimeras for Targeted Protein Stabilization. Nat. Chem. Biol. 2022, 18 (4), 412–421. 10.1038/s41589-022-00971-2. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
