Post-transcriptional mechanisms controlling neurogenesis and direct neuronal reprogramming
- PMID: 38227517
- PMCID: PMC11040297
- DOI: 10.4103/1673-5374.390976
Post-transcriptional mechanisms controlling neurogenesis and direct neuronal reprogramming
Abstract
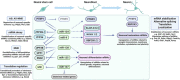
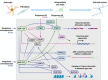
Neurogenesis is a tightly regulated process in time and space both in the developing embryo and in adult neurogenic niches. A drastic change in the transcriptome and proteome of radial glial cells or neural stem cells towards the neuronal state is achieved due to sophisticated mechanisms of epigenetic, transcriptional, and post-transcriptional regulation. Understanding these neurogenic mechanisms is of major importance, not only for shedding light on very complex and crucial developmental processes, but also for the identification of putative reprogramming factors, that harbor hierarchically central regulatory roles in the course of neurogenesis and bare thus the capacity to drive direct reprogramming towards the neuronal fate. The major transcriptional programs that orchestrate the neurogenic process have been the focus of research for many years and key neurogenic transcription factors, as well as repressor complexes, have been identified and employed in direct reprogramming protocols to convert non-neuronal cells, into functional neurons. The post-transcriptional regulation of gene expression during nervous system development has emerged as another important and intricate regulatory layer, strongly contributing to the complexity of the mechanisms controlling neurogenesis and neuronal function. In particular, recent advances are highlighting the importance of specific RNA binding proteins that control major steps of mRNA life cycle during neurogenesis, such as alternative splicing, polyadenylation, stability, and translation. Apart from the RNA binding proteins, microRNAs, a class of small non-coding RNAs that block the translation of their target mRNAs, have also been shown to play crucial roles in all the stages of the neurogenic process, from neural stem/progenitor cell proliferation, neuronal differentiation and migration, to functional maturation. Here, we provide an overview of the most prominent post-transcriptional mechanisms mediated by RNA binding proteins and microRNAs during the neurogenic process, giving particular emphasis on the interplay of specific RNA binding proteins with neurogenic microRNAs. Taking under consideration that the molecular mechanisms of neurogenesis exert high similarity to the ones driving direct neuronal reprogramming, we also discuss the current advances in in vitro and in vivo direct neuronal reprogramming approaches that have employed microRNAs or RNA binding proteins as reprogramming factors, highlighting the so far known mechanisms of their reprogramming action.
Copyright © 2024 Copyright: © 2024 Neural Regeneration Research.
Conflict of interest statement
Figures
Similar articles
-
New insights into the regulatory roles of microRNAs in adult neurogenesis.Curr Opin Pharmacol. 2020 Feb;50:38-45. doi: 10.1016/j.coph.2019.11.003. Epub 2019 Dec 19. Curr Opin Pharmacol. 2020. PMID: 31865131 Review.
-
A miR-124-mediated post-transcriptional mechanism controlling the cell fate switch of astrocytes to induced neurons.Stem Cell Reports. 2023 Apr 11;18(4):915-935. doi: 10.1016/j.stemcr.2023.02.009. Epub 2023 Mar 23. Stem Cell Reports. 2023. PMID: 36963393 Free PMC article.
-
Decoding the ubiquitous role of microRNAs in neurogenesis.Mol Neurobiol. 2017 Apr;54(3):2003-2011. doi: 10.1007/s12035-016-9797-2. Epub 2016 Feb 24. Mol Neurobiol. 2017. PMID: 26910816 Review.
-
Evolving features of human cortical development and the emerging roles of non-coding RNAs in neural progenitor cell diversity and function.Cell Mol Life Sci. 2021 Dec 18;79(1):56. doi: 10.1007/s00018-021-04063-7. Cell Mol Life Sci. 2021. PMID: 34921638 Free PMC article.
-
Controlling the Expression Level of the Neuronal Reprogramming Factors for a Successful Reprogramming Outcome.Cells. 2024 Jul 20;13(14):1223. doi: 10.3390/cells13141223. Cells. 2024. PMID: 39056804 Free PMC article. Review.
Cited by
-
The Hippo effector TEAD1 regulates postnatal murine cerebellar development.Brain Struct Funct. 2025 Mar 10;230(3):42. doi: 10.1007/s00429-025-02903-x. Brain Struct Funct. 2025. PMID: 40064689 Free PMC article.
-
Long-read RNA-sequencing reveals transcript-specific regulation in human-derived cortical neurons.Open Biol. 2025 Jul;15(7):250200. doi: 10.1098/rsob.250200. Epub 2025 Jul 30. Open Biol. 2025. PMID: 40735840 Free PMC article.
-
C-C motif chemokine ligand 2/C-C motif chemokine receptor 2 pathway as a therapeutic target and regulatory mechanism for spinal cord injury.Neural Regen Res. 2025 Aug 1;20(8):2231-2244. doi: 10.4103/NRR.NRR-D-24-00119. Epub 2024 Jul 29. Neural Regen Res. 2025. PMID: 39104168 Free PMC article.
-
Neural stem cell heterogeneity in adult hippocampus.Cell Regen. 2025 Mar 7;14(1):6. doi: 10.1186/s13619-025-00222-4. Cell Regen. 2025. PMID: 40053275 Free PMC article. Review.
References
-
- Abernathy DG, Kim WK, McCoy MJ, Lake AM, Ouwenga R, Lee SW, Xing X, Li D, Lee HJ, Heuckeroth RO, Dougherty JD, Wang T, Yoo AS. MicroRNAs induce a permissive chromatin environment that enables neuronal subtype-specific reprogramming of adult human fibroblasts. Cell Stem Cell. 2017;21:332–348. - PMC - PubMed
LinkOut - more resources
Full Text Sources

