A circRNA ceRNA network involved in cognitive dysfunction after chronic cerebral hypoperfusion
- PMID: 38231472
- PMCID: PMC10866435
- DOI: 10.18632/aging.205387
A circRNA ceRNA network involved in cognitive dysfunction after chronic cerebral hypoperfusion
Abstract
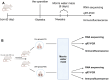
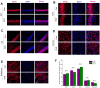
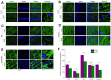
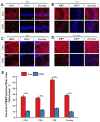
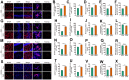
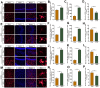
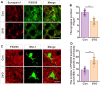
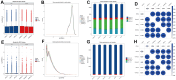
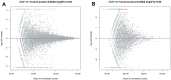
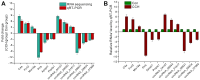
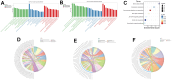
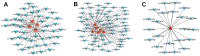
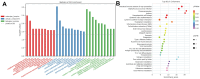
Chronic Cerebral Hypoperfusion (CCH) is associated with cognitive dysfunction, the underlying mechanisms of which remain elusive, hindering the development of effective therapeutic approaches. In this study, we employed an established CCH animal model to delve into neuropathological alterations like oxidative stress, inflammation, neurotransmitter synthesis deficits, and other morphological alterations. Our findings revealed that while the number of neurons remained unchanged, there was a significant reduction in neuronal fibers post-CCH, as evidenced by microtubule-associated protein 2 (MAP2) staining. Moreover, myelin basic protein (MBP) staining showed exacerbated demyelination of neuronal fibers. Furthermore, we observed increased neuroinflammation, proliferation, and activation of astrocytes and microglia, as well as synaptic loss and microglial-mediated synapse engulfment post-CCH. Utilizing RNA sequencing, differential expression analysis displayed alterations in both mRNAs and circRNAs. Following gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses, both showed significant enrichment in immunological and inflammation-related terms and pathways. Importantly, the differentially expressed circular RNAs (DE circRNAs) exhibited a notable coexpression pattern with DE mRNAs. The ternary circRNA-miRNA-mRNA competing endogenous RNAs (ceRNA) network was constructed, and subsequent analysis reiterated the significance of neuroimmunological and neuroinflammatory dysfunction in CCH-induced neuropathological changes and cognitive dysfunction. This study underscores the potential role of circRNAs in these processes, suggesting them as promising therapeutic targets to mitigate the detrimental effects of CCH.
Keywords: RNA sequencing; ceRNA; chronic cerebral hypoperfusion; cognitive dysfunction.
Conflict of interest statement
Figures


Similar articles
-
Identification of Serum Exosome-Derived circRNA-miRNA-TF-mRNA Regulatory Network in Postmenopausal Osteoporosis Using Bioinformatics Analysis and Validation in Peripheral Blood-Derived Mononuclear Cells.Front Endocrinol (Lausanne). 2022 Jun 9;13:899503. doi: 10.3389/fendo.2022.899503. eCollection 2022. Front Endocrinol (Lausanne). 2022. PMID: 35757392 Free PMC article.
-
Comprehensive analysis of the coding and non-coding RNA transcriptome expression profiles of hippocampus tissue in tx-J animal model of Wilson's disease.Sci Rep. 2023 Jun 7;13(1):9252. doi: 10.1038/s41598-023-36503-8. Sci Rep. 2023. PMID: 37286730 Free PMC article.
-
Comparative analysis of circRNA expression profile and circRNA-miRNA-mRNA regulatory network between palmitic and stearic acid-induced lipotoxicity to pancreatic β cells.Bioengineered. 2021 Dec;12(1):9031-9045. doi: 10.1080/21655979.2021.1992333. Bioengineered. 2021. PMID: 34654356 Free PMC article.
-
Identification of a lncRNA/circRNA-miRNA-mRNA ceRNA Network in Alzheimer's Disease.J Integr Neurosci. 2023 Oct 17;22(6):136. doi: 10.31083/j.jin2206136. J Integr Neurosci. 2023. PMID: 38176923
-
Construction and functional enrichment analysis of the competitive endogenous RNA regulatory network for nonarteritic anterior ischemic optic neuropathy based on high-throughput sequencing.Funct Integr Genomics. 2022 Dec;22(6):1253-1267. doi: 10.1007/s10142-022-00914-z. Epub 2022 Nov 10. Funct Integr Genomics. 2022. PMID: 36355332
Cited by
-
Necroptosis in vascular cognitive impairment: mechanisms and therapeutic potential.Front Aging Neurosci. 2025 Jun 25;17:1599773. doi: 10.3389/fnagi.2025.1599773. eCollection 2025. Front Aging Neurosci. 2025. PMID: 40636901 Free PMC article. Review.
References
-
- Duncombe J, Kitamura A, Hase Y, Ihara M, Kalaria RN, Horsburgh K. Chronic cerebral hypoperfusion: a key mechanism leading to vascular cognitive impairment and dementia. Closing the translational gap between rodent models and human vascular cognitive impairment and dementia. Clin Sci (Lond). 2017; 131:2451–68. 10.1042/CS20160727 - DOI - PubMed
-
- Zhao Y, Zhang J, Zheng Y, Zhang Y, Zhang XJ, Wang H, Du Y, Guan J, Wang X, Fu J. NAD+ improves cognitive function and reduces neuroinflammation by ameliorating mitochondrial damage and decreasing ROS production in chronic cerebral hypoperfusion models through Sirt1/PGC-1α pathway. J Neuroinflammation. 2021; 18:207. 10.1186/s12974-021-02250-8 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

