Splicing neoantigen discovery with SNAF reveals shared targets for cancer immunotherapy
- PMID: 38232136
- PMCID: PMC11517820
- DOI: 10.1126/scitranslmed.ade2886
Splicing neoantigen discovery with SNAF reveals shared targets for cancer immunotherapy
Abstract
Immunotherapy has emerged as a crucial strategy to combat cancer by "reprogramming" a patient's own immune system. Although immunotherapy is typically reserved for patients with a high mutational burden, neoantigens produced from posttranscriptional regulation may provide an untapped reservoir of common immunogenic targets for new targeted therapies. To comprehensively define tumor-specific and likely immunogenic neoantigens from patient RNA-Seq, we developed Splicing Neo Antigen Finder (SNAF), an easy-to-use and open-source computational workflow to predict splicing-derived immunogenic MHC-bound peptides (T cell antigen) and unannotated transmembrane proteins with altered extracellular epitopes (B cell antigen). This workflow uses a highly accurate deep learning strategy for immunogenicity prediction (DeepImmuno) in conjunction with new algorithms to rank the tumor specificity of neoantigens (BayesTS) and to predict regulators of mis-splicing (RNA-SPRINT). T cell antigens from SNAF were frequently evidenced as HLA-presented peptides from mass spectrometry (MS) and predict response to immunotherapy in melanoma. Splicing neoantigen burden was attributed to coordinated splicing factor dysregulation. Shared splicing neoantigens were found in up to 90% of patients with melanoma, correlated to overall survival in multiple cancer cohorts, induced T cell reactivity, and were characterized by distinct cells of origin and amino acid preferences. In addition to T cell neoantigens, our B cell focused pipeline (SNAF-B) identified a new class of tumor-specific extracellular neoepitopes, which we termed ExNeoEpitopes. ExNeoEpitope full-length mRNA predictions were tumor specific and were validated using long-read isoform sequencing and in vitro transmembrane localization assays. Therefore, our systematic identification of splicing neoantigens revealed potential shared targets for therapy in heterogeneous cancers.
Conflict of interest statement
COMPETING INTERESTS
The authors declare no competing interests.
Figures





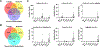

References
-
- Hanahan D, Weinberg RA, Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011). - PubMed
-
- Ott PA, Hu Z, Keskin DB, Shukla SA, Sun J, Bozym DJ, Zhang W, Luoma A, Giobbie-Hurder A, Peter L, Chen C, Olive O, Carter TA, Li S, Lieb DJ, Eisenhaure T, Gjini E, Stevens J, Lane WJ, Javeri I, Nellaiappan K, Salazar AM, Daley H, Seaman M, Buchbinder EI, Yoon CH, Harden M, Lennon N, Gabriel S, Rodig SJ, Barouch DH, Aster JC, Getz G, Wucherpfennig K, Neuberg D, Ritz J, Lander ES, Fritsch EF, Hacohen N, Wu CJ, An immunogenic personal neoantigen vaccine for patients with melanoma. Nature 547, 217–221 (2017). - PMC - PubMed
-
- Hu Z, Leet DE, Allesøe RL, Oliveira G, Li S, Luoma AM, Liu J, Forman J, Huang T, Iorgulescu JB, Holden R, Sarkizova S, Gohil SH, Redd RA, Sun J, Elagina L, Giobbie-Hurder A, Zhang W, Peter L, Ciantra Z, Rodig S, Olive O, Shetty K, Pyrdol J, Uduman M, Lee PC, Bachireddy P, Buchbinder EI, Yoon CH, Neuberg D, Pentelute BL, Hacohen N, Livak KJ, Shukla SA, Olsen LR, Barouch DH, Wucherpfennig KW, Fritsch EF, Keskin DB, Wu CJ, Ott PA, Personal neoantigen vaccines induce persistent memory T cell responses and epitope spreading in patients with melanoma. Nat. Med. 27, 515–525 (2021). - PMC - PubMed
-
- Bauman J, Burris H, Clarke J, Patel M, Cho D, Gutierrez M, Julian R, Scott A, Cohen P, Frederick J, Robert-Tissot C, Zhou H, Mody K, Keating K, Meehan R, Gainor J, 798 Safety, tolerability, and immunogenicity of mRNA-4157 in combination with pembrolizumab in subjects with unresectable solid tumors (KEYNOTE-603): an update. J Immunother Cancer 8 (2020), doi: 10.1136/jitc-2020-SITC2020.0798. - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

