GLS and GOT2 as prognostic biomarkers associated with dendritic cell and immunotherapy response in breast cancer
- PMID: 38234908
- PMCID: PMC10792574
- DOI: 10.1016/j.heliyon.2024.e24163
GLS and GOT2 as prognostic biomarkers associated with dendritic cell and immunotherapy response in breast cancer
Abstract
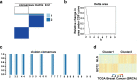
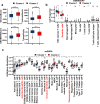
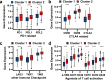
Breast cancer is the females' most common cancer. Targeting the immune microenvironment is a new and promising treatment method for breast cancer. Nevertheless, only a small section of patients can profit by immunotherapy, and improving the ability to accurately predict the potential for immunotherapy response is still awaiting further exploration. In this study, we found that the key factors of glutamine metabolism, glutaminase 1 (GLS) and mitochondrial aspartate transaminase (GOT2), showed opposite expression patterns in breast cancer samples. Based on the expression level of GLS and GOT2, we divided the breast cancer samples into two clusters: Cluster 2 showed GLS expressed higher and GOT2 expressed lower, whereas Cluster 1 showed GOT2 expressed higher and GLS expressed lower. GSEA showed that the clusters were related to pathways of immunity. Further analysis showed that Cluster 2 was positively associated with immunity infiltration. Through WGCNA, we identified a module strongly correlated with glutamine metabolism and immunity and identified 11 dendritic cell-associated genes involved in dendritic cell development, maturation, activation and other functions. In addition, Cluster 2 also showed higher immune checkpoint gene expression, which suggest the Cluster 2 had even better response to immunotherapy. The validation dataset could also be clustered into two groups. Cluster 2 (GLS expressed higher and GOT2 expressed lower) of the validation dataset was also positively associated with dendritic cells and a better immunotherapy response. Thus, these data indicate that GLS and GOT2 are prognostic biomarkers which closely related to dendritic cells and better reacted to immunotherapy in breast cancer.
Keywords: Breast cancer; Dendritic cell; GLS1; GOT2; Immune checkpoint.
© 2024 The Authors.
Conflict of interest statement
The authors declare the following financial interests/personal relationships which may be considered as potential competing interests: Jing Feng reports financial support was provided by 10.13039/501100001809National Natural Science Foundation of China. If there are other authors, they declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures


Similar articles
-
GOT2: New therapeutic target in pancreatic cancer.Genes Dis. 2024 Jul 2;12(4):101370. doi: 10.1016/j.gendis.2024.101370. eCollection 2025 Jul. Genes Dis. 2024. PMID: 40247913 Free PMC article. Review.
-
Preventing BRCA1/ZBRK1 repressor complex binding to the GOT2 promoter results in accelerated aspartate biosynthesis and promotion of cell proliferation.Mol Oncol. 2019 Apr;13(4):959-977. doi: 10.1002/1878-0261.12466. Epub 2019 Mar 1. Mol Oncol. 2019. PMID: 30714292 Free PMC article.
-
Expression of GOT2 Is Epigenetically Regulated by DNA Methylation and Correlates with Immune Infiltrates in Clear-Cell Renal Cell Carcinoma.Curr Issues Mol Biol. 2022 May 25;44(6):2472-2489. doi: 10.3390/cimb44060169. Curr Issues Mol Biol. 2022. PMID: 35735610 Free PMC article.
-
An asparagine metabolism-based classification reveals the metabolic and immune heterogeneity of hepatocellular carcinoma.BMC Med Genomics. 2022 Oct 25;15(1):222. doi: 10.1186/s12920-022-01380-z. BMC Med Genomics. 2022. PMID: 36284275 Free PMC article.
-
Spotlight on GOT2 in Cancer Metabolism.Onco Targets Ther. 2023 Aug 22;16:695-702. doi: 10.2147/OTT.S382161. eCollection 2023. Onco Targets Ther. 2023. PMID: 37635751 Free PMC article. Review.
Cited by
-
Glucose-induced STUB1-GOT2 axis promotes aspartate synthesis and mitochondrial dysfunction in bladder cancer.Cell Death Dis. 2025 Jul 12;16(1):516. doi: 10.1038/s41419-025-07840-5. Cell Death Dis. 2025. PMID: 40651940 Free PMC article.
-
Harnessing amino acid pathways to influence myeloid cell function in tumor immunity.Mol Med. 2025 Feb 4;31(1):44. doi: 10.1186/s10020-025-01099-4. Mol Med. 2025. PMID: 39905317 Free PMC article. Review.
-
GOT2: a moonlighting enzyme at the crossroads of cancer metabolism and theranostics.Front Immunol. 2025 Aug 6;16:1626914. doi: 10.3389/fimmu.2025.1626914. eCollection 2025. Front Immunol. 2025. PMID: 40842990 Free PMC article. Review.
-
GOT2: New therapeutic target in pancreatic cancer.Genes Dis. 2024 Jul 2;12(4):101370. doi: 10.1016/j.gendis.2024.101370. eCollection 2025 Jul. Genes Dis. 2024. PMID: 40247913 Free PMC article. Review.
References
-
- Siegel R.L., et al. Cancer statistics. Ca - Cancer J. Clin. 2022;72(1):7–33. 2022. - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

