A Modular Implementation to Handle and Benchmark Drift Correction for High-Density Extracellular Recordings
- PMID: 38238082
- PMCID: PMC10897502
- DOI: 10.1523/ENEURO.0229-23.2023
A Modular Implementation to Handle and Benchmark Drift Correction for High-Density Extracellular Recordings
Abstract
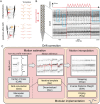
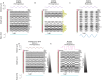
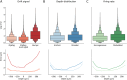
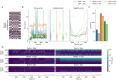
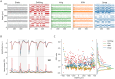
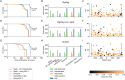
High-density neural devices are now offering the possibility to record from neuronal populations in vivo at unprecedented scale. However, the mechanical drifts often observed in these recordings are currently a major issue for "spike sorting," an essential analysis step to identify the activity of single neurons from extracellular signals. Although several strategies have been proposed to compensate for such drifts, the lack of proper benchmarks makes it hard to assess the quality and effectiveness of motion correction. In this paper, we present a benchmark study to precisely and quantitatively evaluate the performance of several state-of-the-art motion correction algorithms introduced in the literature. Using simulated recordings with induced drifts, we dissect the origins of the errors performed while applying a motion correction algorithm as a preprocessing step in the spike sorting pipeline. We show how important it is to properly estimate the positions of the neurons from extracellular traces in order to correctly estimate the probe motion, compare several interpolation procedures, and highlight what are the current limits for motion correction approaches.
Keywords: benchmark; drift; electrophysiology; ground-truth; neuropixel; spike sorting.
Copyright © 2024 Garcia et al.
Figures

Similar articles
-
MEArec: A Fast and Customizable Testbench Simulator for Ground-truth Extracellular Spiking Activity.Neuroinformatics. 2021 Jan;19(1):185-204. doi: 10.1007/s12021-020-09467-7. Neuroinformatics. 2021. PMID: 32648042 Free PMC article.
-
How Do Spike Collisions Affect Spike Sorting Performance?eNeuro. 2022 Oct 3;9(5):ENEURO.0105-22.2022. doi: 10.1523/ENEURO.0105-22.2022. Print 2022 Sep-Oct. eNeuro. 2022. PMID: 36171060 Free PMC article.
-
SpikeForest, reproducible web-facing ground-truth validation of automated neural spike sorters.Elife. 2020 May 19;9:e55167. doi: 10.7554/eLife.55167. Elife. 2020. PMID: 32427564 Free PMC article.
-
Recent progress in multi-electrode spike sorting methods.J Physiol Paris. 2016 Nov;110(4 Pt A):327-335. doi: 10.1016/j.jphysparis.2017.02.005. Epub 2017 Mar 2. J Physiol Paris. 2016. PMID: 28263793 Free PMC article. Review.
-
Spike sorting algorithms and their efficient hardware implementation: a comprehensive survey.J Neural Eng. 2023 Apr 14;20(2). doi: 10.1088/1741-2552/acc7cc. J Neural Eng. 2023. PMID: 36972585 Review.
Cited by
-
DREDge: robust motion correction for high-density extracellular recordings across species.bioRxiv [Preprint]. 2023 Oct 29:2023.10.24.563768. doi: 10.1101/2023.10.24.563768. bioRxiv. 2023. Update in: Nat Methods. 2025 Apr;22(4):788-800. doi: 10.1038/s41592-025-02614-5. PMID: 37961359 Free PMC article. Updated. Preprint.
-
MEDiCINe: Motion Correction for Neural Electrophysiology Recordings.eNeuro. 2025 Mar 12;12(3):ENEURO.0529-24.2025. doi: 10.1523/ENEURO.0529-24.2025. Print 2025 Mar. eNeuro. 2025. PMID: 39933920 Free PMC article.
-
Is the impact of spontaneous movements on early visual cortex species specific?Trends Neurosci. 2025 Jan;48(1):7-21. doi: 10.1016/j.tins.2024.11.006. Epub 2024 Dec 18. Trends Neurosci. 2025. PMID: 39701910 Review.
-
Structured experience shapes strategy learning and neural dynamics in the medial entorhinal cortex.Res Sq [Preprint]. 2025 May 28:rs.3.rs-6658028. doi: 10.21203/rs.3.rs-6658028/v1. Res Sq. 2025. PMID: 40502753 Free PMC article. Preprint.
-
Structured experience shapes strategy learning and neural dynamics in the medial entorhinal cortex.bioRxiv [Preprint]. 2025 May 13:2025.05.13.653873. doi: 10.1101/2025.05.13.653873. bioRxiv. 2025. PMID: 40463160 Free PMC article. Preprint.
References
-
- Boussard J, Varol E, Dong Lee H, Dethe N, Paninski L (2021) Three-dimensional spike localization and improved motion correction for neuropixels recordings. Adv Neural Inf Process Syst 34:22095–22105.
-
- Boussard J, Windolf C, Hurwitz C, Lee HD, Yu H, Winter O, Paninski L (2023) “DARTsort: A Modular Drift Tracking Spike Sorter for High-Density Multi-electrode Probes.” bioRxiv, pp 2023–08.
MeSH terms
LinkOut - more resources
Full Text Sources
