Impaired activation of plasmacytoid dendritic cells via toll-like receptor 7/9 and STING is mediated by melanoma-derived immunosuppressive cytokines and metabolic drift
- PMID: 38239354
- PMCID: PMC10795195
- DOI: 10.3389/fimmu.2023.1227648
Impaired activation of plasmacytoid dendritic cells via toll-like receptor 7/9 and STING is mediated by melanoma-derived immunosuppressive cytokines and metabolic drift
Abstract
Introduction: Plasmacytoid dendritic cells (pDCs) infiltrate a large set of human cancers. Interferon alpha (IFN-α) produced by pDCs induces growth arrest and apoptosis in tumor cells and modulates innate and adaptive immune cells involved in anti-cancer immunity. Moreover, effector molecules exert tumor cell killing. However, the activation state and clinical relevance of pDCs infiltration in cancer is still largely controversial. In Primary Cutaneous Melanoma (PCM), pDCs density decreases over disease progression and collapses in metastatic melanoma (MM). Moreover, the residual circulating pDC compartment is defective in IFN-α production.
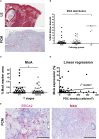
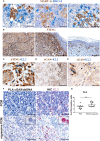
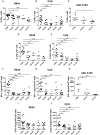
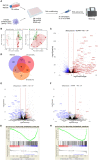
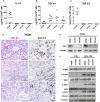
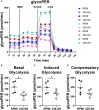
Methods: The activation of tumor-associated pDCs was evaluated by in silico and microscopic analysis. The expression of human myxovirus resistant protein 1 (MxA), as surrogate of IFN-α production, and proximity ligation assay (PLA) to test dsDNA-cGAS activation were performed on human melanoma biopsies. Moreover, IFN-α and CXCL10 production by in vitro stimulated (i.e. with R848, CpG-A, ADU-S100) pDCs exposed to melanoma cell lines supernatants (SN-mel) was tested by intracellular flow cytometry and ELISA. We also performed a bulk RNA-sequencing on SN-mel-exposed pDCs, resting or stimulated with R848. Glycolytic rate assay was performed on SN-mel-exposed pDCs using the Seahorse XFe24 Extracellular Flux Analyzer.
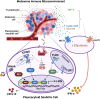
Results: Based on a set of microscopic, functional and in silico analyses, we demonstrated that the melanoma milieu directly impairs IFN-α and CXCL10 production by pDCs via TLR-7/9 and cGAS-STING signaling pathways. Melanoma-derived immunosuppressive cytokines and a metabolic drift represent relevant mechanisms enforcing pDC-mediated melanoma escape.
Discussion: These findings propose a new window of intervention for novel immunotherapy approaches to amplify the antitumor innate immune response in cutaneous melanoma (CM).
Keywords: TGF-β; cGAS-STING; glycolysis; interferon; melanoma; plasmacytoid dendritic cells; toll-like receptor; tumor microenvironment.
Copyright © 2024 Monti, Ferrari, Grosso, Missale, Bugatti, Cancila, Zini, Segala, La Via, Consoli, Orlandi, Valerio, Tripodo, Rossato and Vermi.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The reviewer CM declared a shared affiliation with the authors VG, MO and MR to the handling editor at time of review.
Figures

References
-
- Eggermont A, Blank C, Mandalà M, Long G, Atkinson V, Dalle S, et al. . Adjuvant pembrolizumab versus placebo in resected stage III melanoma (EORTC 1325-MG/KEYNOTE-054): distant metastasis-free survival results from a double-blind, randomised, controlled, phase 3 trial. Lancet Oncol (2021) 22(5):643–54. doi: 10.1016/S1470-2045(21)00065-6 - DOI - PubMed
-
- Larkin J, Weber J, Del Vecchio M, Gogas H, Arance A, Dalle S, et al. . Adjuvant nivolumab versus ipilimumab (CheckMate 238 trial): Reassessment of 4-year efficacy outcomes in patients with stage III melanoma per AJCC-8 staging criteria. Eur J Cancer (2022) 173:285–96. doi: 10.1016/j.ejca.2022.06.041 - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

