Regulatory role of Mss11 in Candida glabrata virulence: adhesion and biofilm formation
- PMID: 38239503
- PMCID: PMC10794409
- DOI: 10.3389/fcimb.2023.1321094
Regulatory role of Mss11 in Candida glabrata virulence: adhesion and biofilm formation
Abstract
Introduction: Candida glabrata has emerged as a fungal pathogen with high infection and mortality rates, and its primary virulence factors are related to adhesion and biofilm formation. These virulence factors in C.glabrata are primarily mediated by epithelial adhesins (Epas), most of which are encoded in subtelomeric regions and regulated by subtelomeric silencing mechanisms. The transcription factor Mss11, known for its regulatory role in adhesion, biofilm formation, and filamentous growth in Saccharomyces cerevisiae and Candida albicans, has also been implicated in the expression of EPA6, suggesting its potential influence on C.glabrata virulence. The present study aims to determine the regulatory role of Mss11 in the virulence of C. glabrata.
Methods: In this work, a Δmss11 null mutant and its complemented strain were constructed from a C.glabrata standard strain. The impact of the transcription factor Mss11 on the virulence of C.glabrata was investigated through a series of phenotypic experiments, including the microbial adhesion to hydrocarbons (MATH) test, adherence assay, biofilm assay, scanning electron microscopy and Galleria mellonella virulence assay. Furthermore, transcriptome sequencing, quantitative reverse transcription polymerase chain reaction (RT-qPCR), and chromatin immunoprecipitation sequencing (ChIP-seq) were employed to investigate the molecular mechanisms behind the regulation of Mss11.
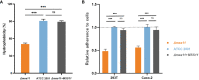
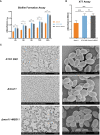
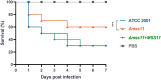
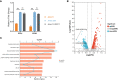
Results: In C.glabrata, the loss of MSS11 led to a significant reduction in several virulence factors including cell surface hydrophobicity, epithelial cell adhesion, and biofilm formation. These observations were consistent with the decreased virulence of the Δmss11 mutant observed in the Galleria mellonella infection model. Further exploration demonstrated that Mss11 modulates C. glabrata virulence by regulating EPA1 and EPA6 expression. It binds to the upstream regions of EPA1 and EPA6, as well as the promoter regions of the subtelomeric silencing-related genes SIR4, RIF1, and RAP1, indicating the dual regulatory role of Mss11.
Conclusion: Mss11 plays a crucial role in C. glabrata adhesion and biofilm formation, and thus has a broad influence on virulence. This regulation is achieved by regulating the expression of EPA1 and EPA6 through both promoter-specific regulation and subtelomeric silencing.
Keywords: Candida glabrata; EPA; Mss11; adhesion; biofilm; subtelomeric silencing; virulence.
Copyright © 2024 Wang, Huang, Zhao, Liu and Xiang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
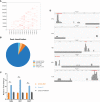
References
-
- Arendrup M. C., Arikan-Akdagli S., Jørgensen K. M., Barac A., Steinmann J., Toscano C., et al. (2023). European candidaemia is characterised by notable differential epidemiology and susceptibility pattern: Results from the ECMM Candida III study. J. Infect. 87, 428–437. doi: 10.1016/j.jinf.2023.08.001 - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

