High UV damage and low repair, but not cytosine deamination, stimulate mutation hotspots at ETS binding sites in melanoma
- PMID: 38241433
- PMCID: PMC10823218
- DOI: 10.1073/pnas.2310854121
High UV damage and low repair, but not cytosine deamination, stimulate mutation hotspots at ETS binding sites in melanoma
Abstract
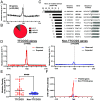
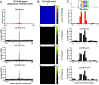
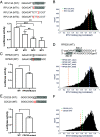
Noncoding mutation hotspots have been identified in melanoma and many of them occur at the binding sites of E26 transformation-specific (ETS) proteins; however, their formation mechanism and functional impacts are not fully understood. Here, we used UV (Ultraviolet) damage sequencing data and analyzed cyclobutane pyrimidine dimer (CPD) formation, DNA repair, and CPD deamination in human cells at single-nucleotide resolution. Our data show prominent CPD hotspots immediately after UV irradiation at ETS binding sites, particularly at sites with a conserved TTCCGG motif, which correlate with mutation hotspots identified in cutaneous melanoma. Additionally, CPDs are repaired slower at ETS binding sites than in flanking DNA. Cytosine deamination in CPDs to uracil is suggested as an important step for UV mutagenesis. However, we found that CPD deamination is significantly suppressed at ETS binding sites, particularly for the CPD hotspot on the 5' side of the ETS motif, arguing against a role for CPD deamination in promoting ETS-associated UV mutations. Finally, we analyzed a subset of frequently mutated promoters, including the ribosomal protein genes RPL13A and RPS20, and found that mutations in the ETS motif can significantly reduce the promoter activity. Thus, our data identify high UV damage and low repair, but not CPD deamination, as the main mechanism for ETS-associated mutations in melanoma and uncover important roles of often-overlooked mutation hotspots in perturbing gene transcription.
Keywords: CPD-seq 2.0; ETS; NER; mutagenesis.
Conflict of interest statement
Competing interests statement:The authors declare no competing interest.
Figures



Similar articles
-
Elevated pyrimidine dimer formation at distinct genomic bases underlies promoter mutation hotspots in UV-exposed cancers.PLoS Genet. 2018 Dec 26;14(12):e1007849. doi: 10.1371/journal.pgen.1007849. eCollection 2018 Dec. PLoS Genet. 2018. PMID: 30586386 Free PMC article.
-
Detecting recurrent passenger mutations in melanoma by targeted UV damage sequencing.Nat Commun. 2023 May 11;14(1):2702. doi: 10.1038/s41467-023-38265-3. Nat Commun. 2023. PMID: 37169747 Free PMC article.
-
ETS transcription factors induce a unique UV damage signature that drives recurrent mutagenesis in melanoma.Nat Commun. 2018 Jul 6;9(1):2626. doi: 10.1038/s41467-018-05064-0. Nat Commun. 2018. PMID: 29980679 Free PMC article.
-
Recurrent Noncoding Mutations in Skin Cancers: UV Damage Susceptibility or Repair Inhibition as Primary Driver?Bioessays. 2019 Mar;41(3):e1800152. doi: 10.1002/bies.201800152. Epub 2019 Feb 25. Bioessays. 2019. PMID: 30801747 Free PMC article. Review.
-
UV wavelength-dependent DNA damage and human non-melanoma and melanoma skin cancer.Photochem Photobiol Sci. 2012 Jan;11(1):90-7. doi: 10.1039/c1pp05144j. Epub 2011 Aug 1. Photochem Photobiol Sci. 2012. PMID: 21804977 Free PMC article. Review.
Cited by
-
Nucleotide excision repair of aflatoxin-induced DNA damage within the 3D human genome organization.Nucleic Acids Res. 2024 Oct 28;52(19):11704-11719. doi: 10.1093/nar/gkae755. Nucleic Acids Res. 2024. PMID: 39258558 Free PMC article.
-
Deciphering the dynamic code: DNA recognition by transcription factors in the ever-changing genome.Transcription. 2024 Jun-Oct;15(3-5):114-138. doi: 10.1080/21541264.2024.2379161. Epub 2024 Jul 20. Transcription. 2024. PMID: 39033307 Free PMC article. Review.
-
Reannotation of cancer mutations based on expressed RNA transcripts reveals functional non-coding mutations in melanoma.Am J Hum Genet. 2025 Jun 5;112(6):1447-1467. doi: 10.1016/j.ajhg.2025.04.005. Epub 2025 May 12. Am J Hum Genet. 2025. PMID: 40359938 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

