Effect of Charge Distribution on the Dynamics of Polyampholytic Disordered Proteins
- PMID: 38250712
- PMCID: PMC10798675
- DOI: 10.1021/acs.macromol.2c01390
Effect of Charge Distribution on the Dynamics of Polyampholytic Disordered Proteins
Abstract
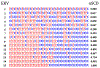
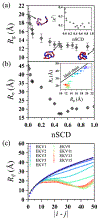
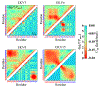
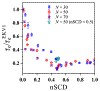
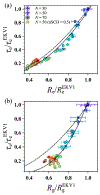
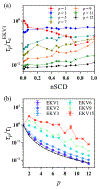
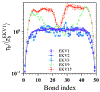
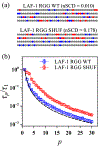
The stability and physiological function of many biomolecular coacervates depend on the structure and dynamics of intrinsically disordered proteins (IDPs) that typically contain a significant fraction of charged residues. Although the effect of relative arrangement of charged residues on IDP conformation is a well-studied problem, the associated changes in dynamics are far less understood. In this work, we systematically interrogate the effects of charge distribution on the chain-level and segmental dynamics of polyampholytic IDPs in dilute solutions. We study a coarse-grained model polyampholyte consisting of an equal fraction of two oppositely charged residues (glutamic acid and lysine) that undergoes a transition from an ideal chain-like conformation for uniformly charge-patterned sequences to a semi-compact conformation for highly charge-segregated sequences. Changes in the chain-level dynamics with increasing charge segregation correlate with changes in conformation. The chain-level and segmental dynamics conform to simple homopolymer models for uniformly charge-patterned sequences but deviate with increasing charge segregation, both in the presence and absence of hydrodynamic interactions. We discuss the significance of these findings, obtained for a model polyampholyte, in the context of a charge-rich intrinsically disordered region of the naturally occurring protein LAF-1. Our findings have important implications for understanding the effects of charge patterning on the dynamics of polyampholytic IDPs in dilute conditions using polymer scaling theories.
Figures

Similar articles
-
Sequence-dependent conformational transitions of disordered proteins during condensation.Chem Sci. 2024 Nov 12;15(47):20056-20063. doi: 10.1039/d4sc05004e. eCollection 2024 Dec 4. Chem Sci. 2024. PMID: 39568948 Free PMC article.
-
Sequence-Dependent Conformational Transitions of Disordered Proteins During Condensation.bioRxiv [Preprint]. 2024 Jan 12:2024.01.11.575294. doi: 10.1101/2024.01.11.575294. bioRxiv. 2024. Update in: Chem Sci. 2024 Nov 12;15(47):20056-20063. doi: 10.1039/d4sc05004e. PMID: 38260590 Free PMC article. Updated. Preprint.
-
Conformations of intrinsically disordered proteins are influenced by linear sequence distributions of oppositely charged residues.Proc Natl Acad Sci U S A. 2013 Aug 13;110(33):13392-7. doi: 10.1073/pnas.1304749110. Epub 2013 Jul 30. Proc Natl Acad Sci U S A. 2013. PMID: 23901099 Free PMC article.
-
Relevance of Electrostatic Charges in Compactness, Aggregation, and Phase Separation of Intrinsically Disordered Proteins.Int J Mol Sci. 2020 Aug 27;21(17):6208. doi: 10.3390/ijms21176208. Int J Mol Sci. 2020. PMID: 32867340 Free PMC article. Review.
-
Spontaneous Switching among Conformational Ensembles in Intrinsically Disordered Proteins.Biomolecules. 2019 Mar 22;9(3):114. doi: 10.3390/biom9030114. Biomolecules. 2019. PMID: 30909517 Free PMC article. Review.
Cited by
-
Equimolar Polyampholyte Hydrogel Synthesis Strategies with Adaptable Properties.Polymers (Basel). 2023 Jul 23;15(14):3131. doi: 10.3390/polym15143131. Polymers (Basel). 2023. PMID: 37514520 Free PMC article.
-
Sequence-dependent material properties of biomolecular condensates and their relation to dilute phase conformations.Nat Commun. 2024 Mar 1;15(1):1912. doi: 10.1038/s41467-024-46223-w. Nat Commun. 2024. PMID: 38429263 Free PMC article.
-
Conformational Properties of Polymers at Droplet Interfaces as Model Systems for Disordered Proteins.ACS Macro Lett. 2023 Nov 21;12(11):1472-1478. doi: 10.1021/acsmacrolett.3c00456. Epub 2023 Oct 19. ACS Macro Lett. 2023. PMID: 37856873 Free PMC article.
-
HYPK: A marginally disordered protein sensitive to charge decoration.Proc Natl Acad Sci U S A. 2024 Apr 30;121(18):e2316408121. doi: 10.1073/pnas.2316408121. Epub 2024 Apr 23. Proc Natl Acad Sci U S A. 2024. PMID: 38657047 Free PMC article.
-
Expanding the molecular language of protein liquid-liquid phase separation.Nat Chem. 2024 Jul;16(7):1113-1124. doi: 10.1038/s41557-024-01489-x. Epub 2024 Mar 29. Nat Chem. 2024. PMID: 38553587 Free PMC article.
References
-
- Brangwynne CP; Eckmann CR; Courson DS; Rybarska A; Hoege C; Gharakhani J; Jülicher F; Hyman AA Germline P Granules Are Liquid Droplets That Localize by Controlled Dissolution/Condensation. Science 2009, 324, 1729–1732. - PubMed
-
- Hammer SK; Avalos JL Harnessing Yeast Organelles for Metabolic Engineering. Nature Chemical Biology 2017, 13, 823–832. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
