Molecular Characterization and Genome Mechanical Features of Two Newly Isolated Polyvalent Bacteriophages Infecting Pseudomonas syringae pv. garcae
- PMID: 38255005
- PMCID: PMC10815195
- DOI: 10.3390/genes15010113
Molecular Characterization and Genome Mechanical Features of Two Newly Isolated Polyvalent Bacteriophages Infecting Pseudomonas syringae pv. garcae
Abstract
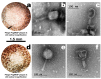
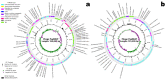
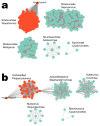
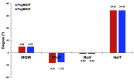
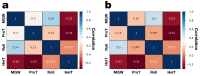
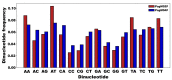
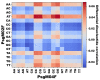
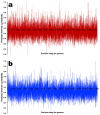
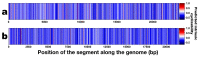
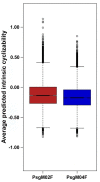
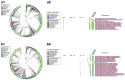
Coffee plants have been targeted by a devastating bacterial disease, a condition known as bacterial blight, caused by the phytopathogen Pseudomonas syringae pv. garcae (Psg). Conventional treatments of coffee plantations affected by the disease involve frequent spraying with copper- and kasugamycin-derived compounds, but they are both highly toxic to the environment and stimulate the appearance of bacterial resistance. Herein, we report the molecular characterization and mechanical features of the genome of two newly isolated (putative polyvalent) lytic phages for Psg. The isolated phages belong to class Caudoviricetes and present a myovirus-like morphotype belonging to the genuses Tequatrovirus (PsgM02F) and Phapecoctavirus (PsgM04F) of the subfamilies Straboviridae (PsgM02F) and Stephanstirmvirinae (PsgM04F), according to recent bacterial viruses' taxonomy, based on their complete genome sequences. The 165,282 bp (PsgM02F) and 151,205 bp (PsgM04F) genomes do not feature any lysogenic-related (integrase) genes and, hence, can safely be assumed to follow a lytic lifestyle. While phage PsgM02F produced a morphogenesis yield of 124 virions per host cell, phage PsgM04F produced only 12 virions per host cell, indicating that they replicate well in Psg with a 50 min latency period. Genome mechanical analyses established a relationship between genome bendability and virion morphogenesis yield within infected host cells.
Keywords: Pseudomonas syringae pv. garcae; Tequatrovirus and Phapecoctavirus genuses; adsorption features; bacteriophage; coffee bacterial blight; genomic structural features; virion morphogenesis yield.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study, in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures


Comment on
-
DNA mechanics and its biological impact.J Mol Biol. 2021 Mar 19;433(6):166861. doi: 10.1016/j.jmb.2021.166861. Epub 2021 Feb 1. J Mol Biol. 2021. PMID: 33539885 Review.
Similar articles
-
Newly isolated phages preying on Pseudomonas syringae pv. garcae: In vitro and ex vivo inactivation studies in coffee plant leafs.Enzyme Microb Technol. 2023 Dec;171:110325. doi: 10.1016/j.enzmictec.2023.110325. Epub 2023 Sep 12. Enzyme Microb Technol. 2023. PMID: 37716050
-
Potential of a newly isolated lytic bacteriophage to control Pseudomonas coronafaciens pv. garcae in coffee plants: Molecular characterization with in vitro and ex vivo experiments.Enzyme Microb Technol. 2025 Mar;184:110573. doi: 10.1016/j.enzmictec.2024.110573. Epub 2024 Dec 15. Enzyme Microb Technol. 2025. PMID: 39700746
-
Control of coffee canker associated with Pseudomonas coronafaciens pv. garcae using a cocktail integrating two virulent polyvalent bacteriophages encapsulated in nanoparticles: in planta studies.J Appl Microbiol. 2025 Jan 6;136(1):lxae310. doi: 10.1093/jambio/lxae310. J Appl Microbiol. 2025. PMID: 39706671
-
Advancements in the Use of Bacteriophages to Combat the Kiwifruit Canker Phytopathogen Pseudomonas syringae pv. actinidiae.Viruses. 2022 Dec 2;14(12):2704. doi: 10.3390/v14122704. Viruses. 2022. PMID: 36560706 Free PMC article. Review.
-
The SPO1-related bacteriophages.Arch Virol. 2010 Oct;155(10):1547-61. doi: 10.1007/s00705-010-0783-0. Epub 2010 Aug 17. Arch Virol. 2010. PMID: 20714761 Review.
References
-
- Ristaino J.B., Anderson P.K., Bebber D.P., Brauman K.A., Cunniffe N.J., Fedoroff N.V., Finegold C., Garrett K.A., Gilligan C.A., Jones C.M., et al. The persistent threat of emerging plant disease pandemics to global food security. Proc. Natl. Acad. Sci. USA. 2021;118:e2022239118. doi: 10.1073/pnas.2022239118. - DOI - PMC - PubMed
-
- Chakraborty S., Newton A.C. Climate change, plant diseases and food security: An overview. Plant Pathol. 2011;60:2–14. doi: 10.1111/j.1365-3059.2010.02411.x. - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

