Favipiravir Analogues as Inhibitors of SARS-CoV-2 RNA-Dependent RNA Polymerase, Combined Quantum Chemical Modeling, Quantitative Structure-Property Relationship, and Molecular Docking Study
- PMID: 38257352
- PMCID: PMC10818557
- DOI: 10.3390/molecules29020441
Favipiravir Analogues as Inhibitors of SARS-CoV-2 RNA-Dependent RNA Polymerase, Combined Quantum Chemical Modeling, Quantitative Structure-Property Relationship, and Molecular Docking Study
Abstract
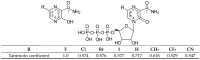
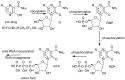
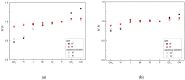
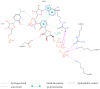
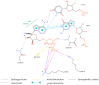
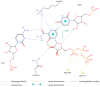
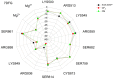
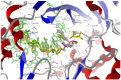
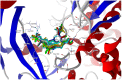
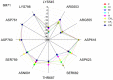
Our study was motivated by the urgent need to develop or improve antivirals for effective therapy targeting RNA viruses. We hypothesized that analogues of favipiravir (FVP), an inhibitor of RNA-dependent RNA polymerase (RdRp), could provide more effective nucleic acid recognition and binding processes while reducing side effects such as cardiotoxicity, hepatotoxicity, teratogenicity, and embryotoxicity. We proposed a set of FVP analogues together with their forms of triphosphate as new SARS-CoV-2 RdRp inhibitors. The main aim of our study was to investigate changes in the mechanism and binding capacity resulting from these modifications. Using three different approaches, QTAIM, QSPR, and MD, the differences in the reactivity, toxicity, binding efficiency, and ability to be incorporated by RdRp were assessed. Two new quantum chemical reactivity descriptors, the relative electro-donating and electro-accepting power, were defined and successfully applied. Moreover, a new quantitative method for comparing binding modes was developed based on mathematical metrics and an atypical radar plot. These methods provide deep insight into the set of desirable properties responsible for inhibiting RdRp, allowing ligands to be conveniently screened. The proposed modification of the FVP structure seems to improve its binding ability and enhance the productive mode of binding. In particular, two of the FVP analogues (the trifluoro- and cyano-) bind very strongly to the RNA template, RNA primer, cofactors, and RdRp, and thus may constitute a very good alternative to FVP.
Keywords: COVID-2019; RNA viruses; SARS-CoV-2; binding modes; favipiravir analogues; hydrogen bonds; interaction patterns.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
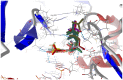


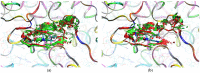


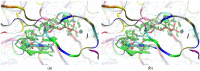



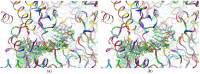
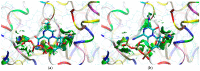

Similar articles
-
Combination of QSAR, molecular docking, molecular dynamic simulation and MM-PBSA: analogues of lopinavir and favipiravir as potential drug candidates against COVID-19.J Biomol Struct Dyn. 2022 May;40(8):3711-3730. doi: 10.1080/07391102.2020.1850355. Epub 2020 Nov 30. J Biomol Struct Dyn. 2022. PMID: 33251975 Free PMC article.
-
State-of-the-Art Molecular Dynamics Simulation Studies of RNA-Dependent RNA Polymerase of SARS-CoV-2.Int J Mol Sci. 2022 Sep 8;23(18):10358. doi: 10.3390/ijms231810358. Int J Mol Sci. 2022. PMID: 36142270 Free PMC article. Review.
-
Determination of Novel SARS-CoV-2 Inhibitors by Combination of Machine Learning and Molecular Modeling Methods.Med Chem. 2024;20(2):153-231. doi: 10.2174/0115734064265609231026063624. Med Chem. 2024. PMID: 37957860
-
Virtual Screening and Quantum Chemistry Analysis for SARS-CoV-2 RNA-Dependent RNA Polymerase Using the ChEMBL Database: Reproduction of the Remdesivir-RTP and Favipiravir-RTP Binding Modes Obtained from Cryo-EM Experiments with High Binding Affinity.Int J Mol Sci. 2022 Sep 20;23(19):11009. doi: 10.3390/ijms231911009. Int J Mol Sci. 2022. PMID: 36232311 Free PMC article.
-
Favipiravir versus other antiviral or standard of care for COVID-19 treatment: a rapid systematic review and meta-analysis.Virol J. 2020 Sep 24;17(1):141. doi: 10.1186/s12985-020-01412-z. Virol J. 2020. PMID: 32972430 Free PMC article.
Cited by
-
Butterfly Effect in Cytarabine: Combined NMR-NQR Experiment, Solid-State Computational Modeling, Quantitative Structure-Property Relationships and Molecular Docking Study.Pharmaceuticals (Basel). 2024 Mar 29;17(4):445. doi: 10.3390/ph17040445. Pharmaceuticals (Basel). 2024. PMID: 38675407 Free PMC article.
-
Exploring pyrazolines as potential inhibitors of NSP3-macrodomain of SARS-CoV-2: synthesis and in silico analysis.Sci Rep. 2025 Jan 4;15(1):767. doi: 10.1038/s41598-024-81711-5. Sci Rep. 2025. PMID: 39755743 Free PMC article.
-
Serine/Threonine Protein Kinases as Attractive Targets for Anti-Cancer Drugs-An Innovative Approach to Ligand Tuning Using Combined Quantum Chemical Calculations, Molecular Docking, Molecular Dynamic Simulations, and Network-like Similarity Graphs.Molecules. 2024 Jul 5;29(13):3199. doi: 10.3390/molecules29133199. Molecules. 2024. PMID: 38999151 Free PMC article.
-
Anti-Butterfly Effect in Ribavirin Studied by Combined Experiment (PXRD/1H-14N NQR Cross-Relaxation Spectroscopy), Quantum Chemical Calculations, Molecular Docking, Molecular Dynamics Simulations, and Novel Structure-Binding Strength and Quadrupolar Indices.Molecules. 2025 Feb 27;30(5):1096. doi: 10.3390/molecules30051096. Molecules. 2025. PMID: 40076317 Free PMC article.
-
The Chameleon Strategy-A Recipe for Effective Ligand Screening for Viral Targets Based on Four Novel Structure-Binding Strength Indices.Viruses. 2024 Jul 3;16(7):1073. doi: 10.3390/v16071073. Viruses. 2024. PMID: 39066235 Free PMC article.
References
-
- WHO Tracking SARS-CoV-2 Variants. [(accessed on 27 September 2023)]. Available online: https://www.who.int/activities/tracking-SARS-CoV-2-variants.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

