Miniaturization of CRISPR/Cas12-Based DNA Sensor Array by Non-Contact Printing
- PMID: 38258263
- PMCID: PMC10818962
- DOI: 10.3390/mi15010144
Miniaturization of CRISPR/Cas12-Based DNA Sensor Array by Non-Contact Printing
Abstract
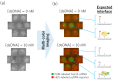
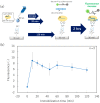
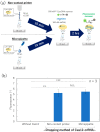
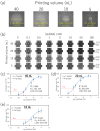
DNA microarrays have been applied for comprehensive genotyping, but remain a drawback in complicated operations. As a solution, we previously reported the solid-phase collateral cleavage (SPCC) system based on the clustered regularly interspaced short palindromic repeat/CRISPR-associated protein 12 (CRISPR/Cas12). Surface-immobilized Cas12-CRISPR RNA (crRNA) can directly hybridize target double-stranded DNA (dsDNA) and subsequently produce a signal via the cleavage of single-stranded DNA (ssDNA) reporter immobilized on the same spot. Therefore, SPCC-based multiplex dsDNA detection can be performed easily. This study reports the miniaturization of SPCC-based spots patterned by a non-contact printer and its performance in comprehensive genotyping on a massively accumulated array. Initially, printing, immobilization, and washing processes of Cas12-crRNA were established to fabricate the non-contact-patterned SPCC-based sensor array. A target dsDNA concentration response was obtained based on the developed sensor array, even with a spot diameter of 0.64 ± 0.05 mm. Also, the limit of detection was 572 pM, 531 pM, and 3.04 nM with 40, 20, and 10 nL-printing of Cas12-crRNA, respectively. Furthermore, the sensor array specifically detected three dsDNA sequences in one-pot multiplexing; therefore, the feasibility of comprehensive genotyping was confirmed. These results demonstrate that our technology can be miniaturized as a CRISPR/Cas12-based microarray by using non-contact printing. In the future, the non-contact-patterned SPCC-based sensor array can be applied as an alternative tool to DNA microarrays.
Keywords: CRISPR/Cas12; accumulated sensor array; collateral cleavage; genotyping; miniaturization; multiplex detection; non-contact bioprinting; printed biosensor; protein array.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures



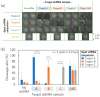
References
-
- Japan: National Action Plan on Antimicrobial Resistance (AMR) [(accessed on 29 November 2023)]. Available online: https://www.who.int/publications/m/item/japan-national-action-plan-on-an...
-
- Kumar A., Roberts D., Wood K.E., Light B., Parrillo J.E., Sharma S., Suppes R., Feinstein D., Zanotti S., Taiberg L., et al. Duration of Hypotension before Initiation of Effective Antimicrobial Therapy Is the Critical Determinant of Survival in Human Septic Shock. Crit. Care Med. 2006;34:1589–1596. doi: 10.1097/01.CCM.0000217961.75225.E9. - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

