This is a preprint.
Structural dynamics of the active HER4 and HER2/HER4 complexes is finely tuned by different growth factors and glycosylation
- PMID: 38260342
- PMCID: PMC10802258
- DOI: 10.1101/2023.10.06.561161
Structural dynamics of the active HER4 and HER2/HER4 complexes is finely tuned by different growth factors and glycosylation
Update in
-
Structural dynamics of the active HER4 and HER2/HER4 complexes is finely tuned by different growth factors and glycosylation.Elife. 2024 Mar 18;12:RP92873. doi: 10.7554/eLife.92873. Elife. 2024. PMID: 38498590 Free PMC article.
Abstract
Human Epidermal growth factor Receptor 4 (HER4 or ERBB4) carries out essential functions in the development and maintenance of the cardiovascular and nervous systems. HER4 activation is regulated by a diverse group of extracellular ligands including the neuregulin (NRG) family and betacellulin (BTC), which promote HER4 homodimerization or heterodimerization with other HER receptors. Important cardiovascular functions of HER4 are exerted via heterodimerization with its close homolog and orphan receptor, HER2. To date structural insights into ligand-mediated HER4 activation have been limited to crystallographic studies of HER4 ectodomain homodimers in complex with NRG1β. Here we report cryo-EM structures of near full-length HER2/HER4 heterodimers and full-length HER4 homodimers bound to NRG1β and BTC. We show that the structures of the heterodimers bound to either ligand are nearly identical and that in both cases the HER2/HER4 heterodimer interface is less dynamic than those observed in structures of HER2/EGFR and HER2/HER3 heterodimers. In contrast, structures of full-length HER4 homodimers bound to NRG1β and BTC display more large-scale dynamics mirroring states previously reported for EGFR homodimers. Our structures also reveal the presence of multiple glycan modifications within HER4 ectodomains, modeled for the first time in HER receptors, that distinctively contribute to the stabilization of HER4 homodimer interfaces over those of HER2/HER4 heterodimers.
Figures

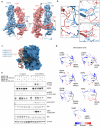


References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
