This is a preprint.
SAX-7/L1CAM acts with the adherens junction proteins MAGI-1, HMR-1/Cadherin, and AFD-1/Afadin to promote glial-mediated dendrite extension
- PMID: 38260503
- PMCID: PMC10802611
- DOI: 10.1101/2024.01.11.575259
SAX-7/L1CAM acts with the adherens junction proteins MAGI-1, HMR-1/Cadherin, and AFD-1/Afadin to promote glial-mediated dendrite extension
Abstract
Adherens junctions (AJs) are a fundamental organizing structure for multicellular life. Although AJs are studied mainly in epithelia, their core function - stabilizing cell contacts by coupling adhesion molecules to the cytoskeleton - is important in diverse tissues. We find that two C. elegans sensory neurons, URX and BAG, require conserved AJ proteins for dendrite morphogenesis. We previously showed that URX and BAG dendrites attach to the embryonic nose via the adhesion molecule SAX-7/L1CAM, acting both in neurons and glia, and then extend by stretch during embryo elongation. Here, we find that a PDZ-binding motif (PB) in the SAX-7 cytoplasmic tail acts with other interaction motifs to promote dendrite extension. Using pull-down assays, we find that the SAX-7 PB binds the multi-PDZ scaffolding protein MAGI-1, which bridges it to the cadherin-catenin complex protein HMP-2/β-catenin. Using cell-specific rescue and depletion, we find that both MAGI-1 and HMR-1/Cadherin act in glia to non-autonomously promote dendrite extension. Double mutant analysis indicates that each protein can act independently of SAX-7, suggesting a multivalent adhesion complex. The SAX-7 PB motif also binds AFD-1/Afadin, loss of which further enhances sax-7 BAG dendrite defects. As MAGI-1, HMR-1, and AFD-1 are all found in epithelial AJs, we propose that an AJ-like complex in glia promotes dendrite extension.
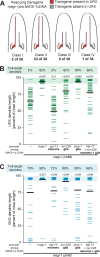
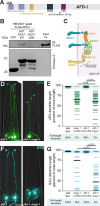
Figures





Similar articles
-
A genome-wide functional screen shows MAGI-1 is an L1CAM-dependent stabilizer of apical junctions in C. elegans.Curr Biol. 2012 Oct 23;22(20):1891-9. doi: 10.1016/j.cub.2012.08.024. Epub 2012 Sep 13. Curr Biol. 2012. PMID: 22981773 Free PMC article.
-
Dendrites with specialized glial attachments develop by retrograde extension using SAX-7 and GRDN-1.Development. 2020 Feb 17;147(4):dev180448. doi: 10.1242/dev.180448. Development. 2020. PMID: 31988188 Free PMC article.
-
Adherens junctions in C. elegans embryonic morphogenesis.Subcell Biochem. 2012;60:279-99. doi: 10.1007/978-94-007-4186-7_12. Subcell Biochem. 2012. PMID: 22674076 Free PMC article. Review.
-
SAX-7/L1CAM and HMR-1/cadherin function redundantly in blastomere compaction and non-muscle myosin accumulation during Caenorhabditis elegans gastrulation.Dev Biol. 2010 Aug 15;344(2):731-44. doi: 10.1016/j.ydbio.2010.05.507. Epub 2010 May 31. Dev Biol. 2010. PMID: 20515680 Free PMC article.
-
Cadherin complexity: recent insights into cadherin superfamily function in C. elegans.Curr Opin Cell Biol. 2012 Oct;24(5):695-701. doi: 10.1016/j.ceb.2012.06.008. Epub 2012 Jul 19. Curr Opin Cell Biol. 2012. PMID: 22819515 Free PMC article. Review.
References
-
- Adle-Biassette H., Saugier-Veber P., Fallet-Bianco C., Delezoide A.-L., Razavi F., Drouot N., Bazin A., Beaufrère A.-M., Bessières B., Blesson S., et al. (2013). Neuropathological review of 138 cases genetically tested for X-linked hydrocephalus: evidence for closely related clinical entities of unknown molecular bases. Acta Neuropathol 126, 427–442. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
