PIGB maintains nuclear lamina organization in skeletal muscle of Drosophila
- PMID: 38261271
- PMCID: PMC10808031
- DOI: 10.1083/jcb.202301062
PIGB maintains nuclear lamina organization in skeletal muscle of Drosophila
Abstract
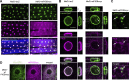
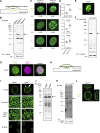
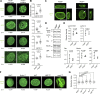
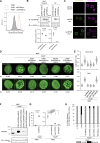
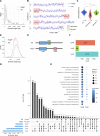
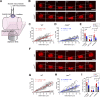
The nuclear lamina (NL) plays various roles and participates in nuclear integrity, chromatin organization, and transcriptional regulation. Lamin proteins, the main components of the NL, form a homogeneous meshwork structure under the nuclear envelope. Lamins are essential, but it is unknown whether their homogeneous distribution is important for nuclear function. Here, we found that PIGB, an enzyme involved in glycosylphosphatidylinositol (GPI) synthesis, is responsible for the homogeneous lamin meshwork in Drosophila. Loss of PIGB resulted in heterogeneous distributions of B-type lamin and lamin-binding proteins in larval muscles. These phenotypes were rescued by expression of PIGB lacking GPI synthesis activity. The PIGB mutant exhibited changes in lamina-associated domains that are large heterochromatic genomic regions in the NL, reduction of nuclear stiffness, and deformation of muscle fibers. These results suggest that PIGB maintains the homogeneous meshwork of the NL, which may be essential for chromatin distribution and nuclear mechanical properties.
© 2024 Yamamoto-Hino et al.
Conflict of interest statement
Disclosures: The authors declare no competing interests exist.
Figures









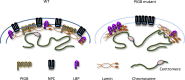
Similar articles
-
Nuclear envelope remodelling during human spermiogenesis involves somatic B-type lamins and a spermatid-specific B3 lamin isoform.Mol Hum Reprod. 2015 Mar;21(3):225-36. doi: 10.1093/molehr/gau111. Epub 2014 Dec 4. Mol Hum Reprod. 2015. PMID: 25477337
-
Lamin is essential for nuclear localization of the GPI synthesis enzyme PIG-B and GPI-anchored protein production in Drosophila.J Cell Sci. 2020 Mar 26;133(6):jcs238527. doi: 10.1242/jcs.238527. J Cell Sci. 2020. PMID: 32051283 Free PMC article.
-
Non-farnesylated B-type lamin can tether chromatin inside the nucleus and its chromatin interaction requires the Ig-fold region.Chromosoma. 2017 Feb;126(1):125-144. doi: 10.1007/s00412-016-0581-x. Epub 2016 Feb 19. Chromosoma. 2017. PMID: 26892013
-
Laminopathies: what can humans learn from fruit flies.Cell Mol Biol Lett. 2018 Jul 6;23:32. doi: 10.1186/s11658-018-0093-1. eCollection 2018. Cell Mol Biol Lett. 2018. PMID: 30002683 Free PMC article. Review.
-
Lamins: The backbone of the nucleocytoskeleton interface.Curr Opin Cell Biol. 2024 Feb;86:102313. doi: 10.1016/j.ceb.2023.102313. Epub 2024 Jan 22. Curr Opin Cell Biol. 2024. PMID: 38262116 Review.
Cited by
-
Nuclear Structure, Size Regulation, and Role in Cell Migration.Cells. 2024 Dec 23;13(24):2130. doi: 10.3390/cells13242130. Cells. 2024. PMID: 39768219 Free PMC article. Review.
-
Sphingomyelin regulates the transcriptional machinery in nuclear lipid microdomains.Commun Biol. 2025 Aug 29;8(1):1303. doi: 10.1038/s42003-025-08697-2. Commun Biol. 2025. PMID: 40883422 Free PMC article.
References
-
- Ashery-Padan, R., Ulitzur N., Arbel A., Goldberg M., Weiss A.M., Maus N., Fisher P.A., and Gruenbaum Y.. 1997. Localization and posttranslational modifications of otefin, a protein required for vesicle attachment to chromatin, during Drosophila melanogaster development. Mol. Cell. Biol. 17:4114–4123. 10.1128/MCB.17.7.4114 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

