Yeast 26S proteasome nuclear import is coupled to nucleus-specific degradation of the karyopherin adaptor protein Sts1
- PMID: 38267508
- PMCID: PMC10808114
- DOI: 10.1038/s41598-024-52352-5
Yeast 26S proteasome nuclear import is coupled to nucleus-specific degradation of the karyopherin adaptor protein Sts1
Abstract
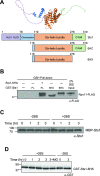
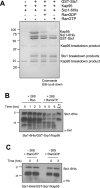
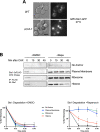
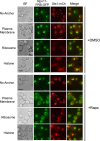
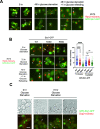
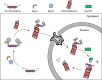
In eukaryotes, the ubiquitin-proteasome system is an essential pathway for protein degradation and cellular homeostasis. 26S proteasomes concentrate in the nucleus of budding yeast Saccharomyces cerevisiae due to the essential import adaptor protein Sts1 and the karyopherin-α protein Srp1. Here, we show that Sts1 facilitates proteasome nuclear import by recruiting proteasomes to the karyopherin-α/β heterodimer. Following nuclear transport, the karyopherin proteins are likely separated from Sts1 through interaction with RanGTP in the nucleus. RanGTP-induced release of Sts1 from the karyopherin proteins initiates Sts1 proteasomal degradation in vitro. Sts1 undergoes karyopherin-mediated nuclear import in the absence of proteasome interaction, but Sts1 degradation in vivo is only observed when proteasomes successfully localize to the nucleus. Sts1 appears to function as a proteasome import factor during exponential growth only, as it is not found in proteasome storage granules (PSGs) during prolonged glucose starvation, nor does it appear to contribute to the rapid nuclear reimport of proteasomes following glucose refeeding and PSG dissipation. We propose that Sts1 acts as a single-turnover proteasome nuclear import factor by recruiting karyopherins for transport and undergoing subsequent RanGTP-initiated ubiquitin-independent proteasomal degradation in the nucleus.
© 2024. The Author(s).
Conflict of interest statement
No competing interests declared.
Figures
Similar articles
-
The Sts1 nuclear import adapter uses a non-canonical bipartite nuclear localization signal and is directly degraded by the proteasome.J Cell Sci. 2020 Mar 19;133(6):jcs236158. doi: 10.1242/jcs.236158. J Cell Sci. 2020. PMID: 32041904 Free PMC article.
-
Nuclear import factor Srp1 and its associated protein Sts1 couple ribosome-bound nascent polypeptides to proteasomes for cotranslational degradation.J Biol Chem. 2014 Jan 31;289(5):2701-10. doi: 10.1074/jbc.M113.524926. Epub 2013 Dec 12. J Biol Chem. 2014. PMID: 24338021 Free PMC article.
-
Sts1 plays a key role in targeting proteasomes to the nucleus.J Biol Chem. 2011 Jan 28;286(4):3104-18. doi: 10.1074/jbc.M110.135863. Epub 2010 Nov 12. J Biol Chem. 2011. PMID: 21075847 Free PMC article.
-
Nuclear transport of yeast proteasomes.Biomolecules. 2014 Oct 20;4(4):940-55. doi: 10.3390/biom4040940. Biomolecules. 2014. PMID: 25333764 Free PMC article. Review.
-
Intracellular localization of the proteasome in response to stress conditions.J Biol Chem. 2022 Jul;298(7):102083. doi: 10.1016/j.jbc.2022.102083. Epub 2022 May 27. J Biol Chem. 2022. PMID: 35636514 Free PMC article. Review.
Cited by
-
Limiting 20S proteasome assembly leads to unbalanced nucleo-cytoplasmic distribution of 26S/30S proteasomes and chronic proteotoxicity.iScience. 2024 Oct 4;27(11):111095. doi: 10.1016/j.isci.2024.111095. eCollection 2024 Nov 15. iScience. 2024. PMID: 39473973 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

