KAS-seq profiling captures transcription dynamics during oocyte maturation
- PMID: 38267939
- PMCID: PMC10807090
- DOI: 10.1186/s13048-023-01342-8
KAS-seq profiling captures transcription dynamics during oocyte maturation
Abstract
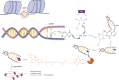
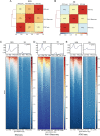
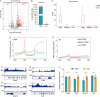
In fully grown oocytes, the genome is considered to be globally transcriptionally silenced. However, this conclusion is primarily derived from the results obtained through immunofluorescence staining or inferred from the highly condensed state of chromosomes, lacking more direct evidence. Here, by using a kethoxal-assisted single-stranded DNA sequencing (KAS-seq) approach, we investigated the landscape of single-strand DNA (ssDNA) throughout the genome and provided a readout of the activity and dynamics of transcription during oocyte meiotic maturation. In non-surrounded nucleolus (NSN) oocytes, we observed a robust KAS-seq signal, indicating the high transcriptional activity. In surrounded nucleolus (SN) oocytes, the presence of ssDNA still persists although the KAS-seq signal was relatively weak, suggesting the presence of transcription. Accompanying with the meiotic resumption, the transcriptional activity gradually decreased, and global repression was detected in matured oocytes. Moreover, we preformed the integrative genomics analysis to dissect the transcriptional dynamics during mouse oocyte maturation. In sum, the present study delineates the detailed transcriptional activity during mammalian oocyte maturation.
Keywords: KAS-seq; Meiosis; Oocytes; Transcription; ssDNA.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
Maternal factors required for oocyte developmental competence in mice: transcriptome analysis of non-surrounded nucleolus (NSN) and surrounded nucleolus (SN) oocytes.Cell Cycle. 2013 Jun 15;12(12):1928-38. doi: 10.4161/cc.24991. Epub 2013 May 13. Cell Cycle. 2013. PMID: 23673344 Free PMC article.
-
Transcriptome and DNA methylation profiling during the NSN to SN transition in mouse oocytes.BMC Mol Cell Biol. 2025 Jan 3;26(1):2. doi: 10.1186/s12860-024-00527-3. BMC Mol Cell Biol. 2025. PMID: 39754059 Free PMC article.
-
Transcriptional activity associated with meiotic competence in fully grown mouse GV oocytes.Zygote. 2002 Nov;10(4):327-32. doi: 10.1017/s0967199402004069. Zygote. 2002. PMID: 12463528
-
KAS-seq: genome-wide sequencing of single-stranded DNA by N3-kethoxal-assisted labeling.Nat Protoc. 2022 Feb;17(2):402-420. doi: 10.1038/s41596-021-00647-6. Epub 2022 Jan 10. Nat Protoc. 2022. PMID: 35013616 Free PMC article. Review.
-
Chromatin Configuration in Diplotene Mouse and Human Oocytes during the Period of Transcriptional Activity Extinction.Int J Mol Sci. 2023 Jul 15;24(14):11517. doi: 10.3390/ijms241411517. Int J Mol Sci. 2023. PMID: 37511273 Free PMC article. Review.
Cited by
-
Integrated Multi-Omics Analysis Reveals Key Regulators of Bovine Oocyte Maturation.Int J Mol Sci. 2025 Apr 23;26(9):3973. doi: 10.3390/ijms26093973. Int J Mol Sci. 2025. PMID: 40362214 Free PMC article.
-
Rescue in vitro maturation using ovarian support cells of human oocytes from conventional stimulation cycles yields oocytes with improved nuclear maturation and transcriptomic resemblance to in vivo matured oocytes.J Assist Reprod Genet. 2024 Aug;41(8):2021-2036. doi: 10.1007/s10815-024-03143-4. Epub 2024 May 30. J Assist Reprod Genet. 2024. PMID: 38814543 Free PMC article.
-
The chromatin accessibility landscape of mouse oocytes during configuration transition.Cell Prolif. 2025 Jan;58(1):e13733. doi: 10.1111/cpr.13733. Epub 2024 Sep 8. Cell Prolif. 2025. PMID: 39245646 Free PMC article.
-
Single-cell multiomic analysis reveals methylome and transcriptome deviations following oocyte maturation in vitro.Reproduction. 2025 Jul 10;170(2):e250011. doi: 10.1530/REP-25-0011. Print 2025 Aug 1. Reproduction. 2025. PMID: 40569608 Free PMC article.
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

