RNA-Seq of Phenotypically Distinct Eimeria maxima Strains Reveals Coordinated and Contrasting Maturation and Shared Sporogonic Biomarkers with Eimeria acervulina
- PMID: 38276148
- PMCID: PMC10818985
- DOI: 10.3390/pathogens13010002
RNA-Seq of Phenotypically Distinct Eimeria maxima Strains Reveals Coordinated and Contrasting Maturation and Shared Sporogonic Biomarkers with Eimeria acervulina
Abstract
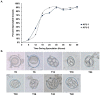
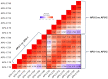
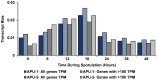
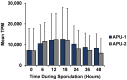
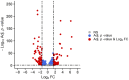
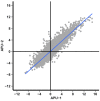
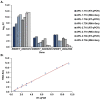
Strains of Eimeria maxima, an enteric parasite of poultry, vary in virulence. Here, we performed microscopy and RNA sequencing on oocysts of strains APU-1 (which exhibits more virulence) and APU-2. Although each underwent parallel development, APU-1 initially approached maturation more slowly. Each strain sporulated by hour 36; their gene expression diverged somewhat thereafter. Candidate biomarkers of viability included 58 genes contributing at least 1000 Transcripts Per Million throughout sporulation, such as cation-transporting ATPases and zinc finger domain-containing proteins. Many genes resemble constitutively expressed genes also important to Eimeria acervulina. Throughout sporulation, the expression of only a few genes differed between strains; these included cyclophilin A, EF-1α, and surface antigens (SAGs). Mature and immature oocysts uniquely differentially express certain genes, such as an X-Pro dipeptidyl-peptidase domain-containing protein in immature oocysts and a profilin in mature oocysts. The immature oocysts of each strain expressed more phosphoserine aminotransferase and the mature oocysts expressed more SAGs and microneme proteins. These data illuminate processes influencing sporulation in Eimeria and related genera, such as Cyclospora, and identify biological processes which may differentiate them. Drivers of development and senescence may provide tools to assess the viability of oocysts, which would greatly benefit the poultry industry and food safety applications.
Keywords: Cyclospora; Eimeria; coccidia; control; development; differential expression; oocyst; sporulation; transcription; vaccine.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Dynamically expressed genes provide candidate viability biomarkers in a model coccidian.PLoS One. 2021 Oct 1;16(10):e0258157. doi: 10.1371/journal.pone.0258157. eCollection 2021. PLoS One. 2021. PMID: 34597342 Free PMC article.
-
UV radiation at 222, 254, and 282 nm inhibits sporulation and suppresses infectivity of Eimeria acervulina oocysts.Microbiol Spectr. 2025 Mar 4;13(3):e0243924. doi: 10.1128/spectrum.02439-24. Epub 2025 Feb 6. Microbiol Spectr. 2025. PMID: 39912693 Free PMC article.
-
Differing susceptibilities of Eimeria acervulina, Eimeria maxima, and Eimeria tenella oocysts to desiccation.J Parasitol. 2013 Oct;99(5):899-902. doi: 10.1645/13-192.1. Epub 2013 Apr 25. J Parasitol. 2013. PMID: 23617755
-
Epidemiological aspects of the use of live anticoccidial vaccines for chickens.Int J Parasitol. 1998 Jul;28(7):1089-98. doi: 10.1016/s0020-7519(98)00066-6. Int J Parasitol. 1998. PMID: 9724880 Review.
-
Eimeria proteins: order amidst disorder.Parasit Vectors. 2022 Jan 24;15(1):38. doi: 10.1186/s13071-022-05159-0. Parasit Vectors. 2022. PMID: 35073987 Free PMC article. Review.
Cited by
-
Intraspecific Variations in Biology and Pathogenesis of Two Eimeria maxima Isolates From Distinct Geographic Locations.Vet Med Sci. 2025 May;11(3):e70235. doi: 10.1002/vms3.70235. Vet Med Sci. 2025. PMID: 40207378 Free PMC article.
-
Molecular characterization of cDNA coding for 33.5 and 41 kDa oocyst and sporocyst proteins that are differentially regulated in different strains of Eimeria maxima.Front Vet Sci. 2024 Sep 27;11:1445646. doi: 10.3389/fvets.2024.1445646. eCollection 2024. Front Vet Sci. 2024. PMID: 39397810 Free PMC article.
-
Development of a TaqMan polymerase chain reaction detection method for the precise identification and quantification of an attenuated Eimeria maxima vaccine strain in poultry.Front Vet Sci. 2024 May 22;11:1397166. doi: 10.3389/fvets.2024.1397166. eCollection 2024. Front Vet Sci. 2024. PMID: 38840634 Free PMC article.
References
-
- Stelzer S., Basso W., Benavides Silvan J., Ortega-Mora L.M., Maksimov P., Gethmann J., Conraths F.J., Schares G. Toxoplasma gondii infection and toxoplasmosis in farm animals: Risk factors and economic impact. Food Waterborne Parasitol. 2019;15:e00037. doi: 10.1016/j.fawpar.2019.e00037. - DOI - PMC - PubMed
-
- Shivaramaiah C., Barta J.R., Hernandez-Velasco X., Tellez G., Hargis B.M. Coccidiosis: Recent advancements in the immunobiology of Eimeria species, preventive measures, and the importance of vaccination as a control tool against these Apicomplexan parasites. Vet. Med. 2014;5:23–34. doi: 10.2147/VMRR.S57839. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

