Expanding the Genetic Code of Xenopus laevis Embryos
- PMID: 38277773
- PMCID: PMC10877573
- DOI: 10.1021/acschembio.3c00686
Expanding the Genetic Code of Xenopus laevis Embryos
Abstract
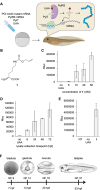
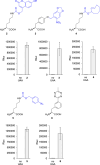
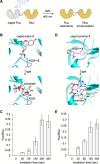
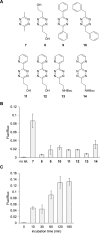
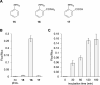
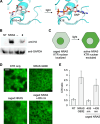
The incorporation of unnatural amino acids into proteins through genetic code expansion has been successfully adapted to African claw-toed frog embryos. Six unique unnatural amino acids are incorporated site-specifically into proteins and demonstrate robust and reliable protein expression. Of these amino acids, several are caged analogues that can be used to establish conditional control over enzymatic activity. Using light or small molecule triggers, we exhibit activation and tunability of protein functions in live embryos. This approach was then applied to optical control over the activity of a RASopathy mutant of NRAS, taking advantage of generating explant cultures from Xenopus. Taken together, genetic code expansion is a robust approach in the Xenopus model to incorporate novel chemical functionalities into proteins of interest to study their function and role in a complex biological setting.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
Similar articles
-
Expanding the genetic code in Xenopus laevis oocytes.Chembiochem. 2013 Jan 21;14(2):230-5. doi: 10.1002/cbic.201200515. Epub 2013 Jan 4. Chembiochem. 2013. PMID: 23292655
-
Using genetically incorporated unnatural amino acids to control protein functions in mammalian cells.Essays Biochem. 2019 Jul 3;63(2):237-266. doi: 10.1042/EBC20180042. Print 2019 Jul 3. Essays Biochem. 2019. PMID: 31092687 Free PMC article. Review.
-
Expanding the Genetic Code for Neuronal Studies.Chembiochem. 2020 Nov 16;21(22):3169-3179. doi: 10.1002/cbic.202000300. Epub 2020 Jul 20. Chembiochem. 2020. PMID: 32531101 Free PMC article. Review.
-
Genetic Code Expansion of Mammalian Cells with Unnatural Amino Acids.Curr Protoc Chem Biol. 2015 Sep 1;7(3):187-199. doi: 10.1002/9780470559277.ch150038. Curr Protoc Chem Biol. 2015. PMID: 26331526
-
A Simplified Protocol to Incorporate the Fluorescent Unnatural Amino Acid ANAP into Xenopus laevis Oocyte-Expressed P2X7 Receptors.Methods Mol Biol. 2022;2510:193-216. doi: 10.1007/978-1-0716-2384-8_10. Methods Mol Biol. 2022. PMID: 35776326
Cited by
-
Optogenetics with Atomic Precision─A Comprehensive Review of Optical Control of Protein Function through Genetic Code Expansion.Chem Rev. 2025 Feb 26;125(4):1663-1717. doi: 10.1021/acs.chemrev.4c00224. Epub 2025 Feb 10. Chem Rev. 2025. PMID: 39928721 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

