Functional characterization of calmodulin-like proteins, CML13 and CML14, as novel light chains of Arabidopsis class VIII myosins
- PMID: 38280207
- PMCID: PMC11272076
- DOI: 10.1093/jxb/erae031
Functional characterization of calmodulin-like proteins, CML13 and CML14, as novel light chains of Arabidopsis class VIII myosins
Abstract
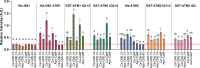
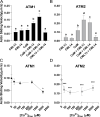
Myosins are important motor proteins that associate with the actin cytoskeleton. Structurally, myosins function as heteromeric complexes where smaller light chains, such as calmodulin (CaM), bind to isoleucine-glutamine (IQ) domains in the neck region to facilitate mechano-enzymatic activity. We recently identified Arabidopsis CaM-like (CML) proteins CML13 and CML14 as interactors of proteins containing multiple IQ domains, including a myosin VIII. Here, we demonstrate that CaM, CML13, and CML14 bind the neck region of all four Arabidopsis myosin VIII isoforms. Among CMLs tested for binding to myosins VIIIs, CaM, CML13, and CML14 gave the strongest signals using in planta split-luciferase protein interaction assays. In vitro, recombinant CaM, CML13, and CML14 showed specific, high-affinity, calcium-independent binding to the IQ domains of myosin VIIIs. CaM, CML13, and CML14 co-localized to plasma membrane-bound puncta when co-expressed with red fluorescent protein-myosin fusion proteins containing IQ and tail domains of myosin VIIIs. In vitro actin motility assays using recombinant myosin VIIIs demonstrated that CaM, CML13, and CML14 function as light chains. Suppression of CML13 or CML14 expression using RNA silencing resulted in a shortened-hypocotyl phenotype, similar to that observed in a quadruple myosin mutant, myosin viii4KO. Collectively, our data indicate that Arabidopsis CML13 and CML14 are novel myosin VIII light chains.
Keywords: Calmodulin-like proteins; cytoskeleton; motor protein; myosin light chains; plant myosins.
© The Author(s) 2024. Published by Oxford University Press on behalf of the Society for Experimental Biology. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Conflict of interest statement
The authors have no conflicts to declare.
Figures







Similar articles
-
Arabidopsis calmodulin-like proteins CML13 and CML14 interact with proteins that have IQ domains.Plant Cell Environ. 2023 Aug;46(8):2470-2491. doi: 10.1111/pce.14616. Epub 2023 May 24. Plant Cell Environ. 2023. PMID: 37222394
-
Arabidopsis Calmodulin-like Proteins CML13 and CML14 Interact with Calmodulin-Binding Transcriptional Activators and Function in Salinity Stress Response.Plant Cell Physiol. 2024 Feb 15;65(2):282-300. doi: 10.1093/pcp/pcad152. Plant Cell Physiol. 2024. PMID: 38036467
-
Arabidopsis CML13 and CML14 Have Essential and Overlapping Roles in Plant Development.Plant Cell Physiol. 2024 Feb 15;65(2):228-242. doi: 10.1093/pcp/pcad142. Plant Cell Physiol. 2024. PMID: 37946525
-
[Calcium ions in the regulation of acto-myosin interactions].Postepy Biochem. 2012;58(4):437-51. Postepy Biochem. 2012. PMID: 23662437 Review. Polish.
-
The Ayerst Award Lecture 1990. Calcium-dependent mechanisms of regulation of smooth muscle contraction.Biochem Cell Biol. 1991 Dec;69(12):771-800. doi: 10.1139/o91-119. Biochem Cell Biol. 1991. PMID: 1818584 Review.
Cited by
-
Characterization of the Calmodulin-Like Protein Family in Chara braunii and their Conserved Interaction with the Calmodulin-Binding Transcription Activator Family.Plant Cell Physiol. 2024 Dec 21;65(12):2040-2053. doi: 10.1093/pcp/pcae127. Plant Cell Physiol. 2024. PMID: 39460541 Free PMC article.
-
The calcium sensor AtCML8 contributes to Arabidopsis plant cell growth by modulating the brassinosteroid signaling pathway.Plant J. 2025 Jan;121(1):e17179. doi: 10.1111/tpj.17179. Epub 2024 Dec 2. Plant J. 2025. PMID: 39621453 Free PMC article.
-
Transcriptome Analysis of the CML Gene Family in Bletilla striata and Regulation of Militarine Synthesis Under Sodium Acetate and Salicylic Acid Treatments.Plants (Basel). 2025 Mar 28;14(7):1052. doi: 10.3390/plants14071052. Plants (Basel). 2025. PMID: 40219120 Free PMC article.
References
-
- Abu-Abied M, Golomb L, Belausov E, Huang S, Geiger B, Kam Z, Staiger CJ, Sadot E. 2006. Identification of plant cytoskeleton-interacting proteins by screening for actin stress fiber association in mammalian fibroblasts. The Plant Journal 48, 367–379. - PubMed
-
- Alaimo A, Malo C, Areso P, Aloria K, Millet O, Villarroel A. 2013. The use of dansyl-calmodulin to study interactions with channels and other proteins. Methods in Molecular Biology 998, 217–231. - PubMed
-
- Altman D. 2013. Myosin: fundamental properties and structure. In: Roberts GCK, ed. Encyclopaedia of biophysics. Berlin, Heidelberg: Springer, 1664–1671.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

