MIWE: detecting the critical states of complex biological systems by the mutual information weighted entropy
- PMID: 38280998
- PMCID: PMC10822190
- DOI: 10.1186/s12859-024-05667-z
MIWE: detecting the critical states of complex biological systems by the mutual information weighted entropy
Abstract
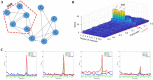
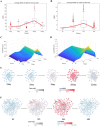
Complex biological systems often undergo sudden qualitative changes during their dynamic evolution. These critical transitions are typically characterized by a catastrophic progression of the system. Identifying the critical point is critical to uncovering the underlying mechanisms of complex biological systems. However, the system may exhibit minimal changes in its state until the critical point is reached, and in the face of high throughput and strong noise data, traditional biomarkers may not be effective in distinguishing the critical state. In this study, we propose a novel approach, mutual information weighted entropy (MIWE), which uses mutual information between genes to build networks and identifies critical states by quantifying molecular dynamic differences at each stage through weighted differential entropy. The method is applied to one numerical simulation dataset and four real datasets, including bulk and single-cell expression datasets. The critical states of the system can be recognized and the robustness of MIWE method is verified by numerical simulation under the influence of different noises. Moreover, we identify two key transcription factors (TFs), CREB1 and CREB3, that regulate downstream signaling genes to coordinate cell fate commitment. The dark genes in the single-cell expression datasets are mined to reveal the potential pathway regulation mechanism.
Keywords: Critical state; Differential entropy; Dynamic network biomarker (DNB); Mutual information.
© 2024. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures





Similar articles
-
CPMI: comprehensive neighborhood-based perturbed mutual information for identifying critical states of complex biological processes.BMC Bioinformatics. 2024 Jun 15;25(1):215. doi: 10.1186/s12859-024-05836-0. BMC Bioinformatics. 2024. PMID: 38879513 Free PMC article.
-
SPNE: sample-perturbed network entropy for revealing critical states of complex biological systems.Brief Bioinform. 2023 Mar 19;24(2):bbad028. doi: 10.1093/bib/bbad028. Brief Bioinform. 2023. PMID: 36705581
-
mNFE: microbiome network flow entropy for detecting pre-disease states of type 1 diabetes.Gut Microbes. 2024 Jan-Dec;16(1):2327349. doi: 10.1080/19490976.2024.2327349. Epub 2024 Mar 21. Gut Microbes. 2024. PMID: 38512768 Free PMC article.
-
Information processing in bacteria: memory, computation, and statistical physics: a key issues review.Rep Prog Phys. 2016 May;79(5):052601. doi: 10.1088/0034-4885/79/5/052601. Epub 2016 Apr 8. Rep Prog Phys. 2016. PMID: 27058315 Free PMC article. Review.
-
Development of a dynamic network biomarkers method and its application for detecting the tipping point of prior disease development.Comput Struct Biotechnol J. 2022 Feb 24;20:1189-1197. doi: 10.1016/j.csbj.2022.02.019. eCollection 2022. Comput Struct Biotechnol J. 2022. PMID: 35317238 Free PMC article. Review.
Cited by
-
Revealing the critical state and identifying individualized dynamic network biomarker for type 2 diabetes through advanced analysis methods on individual basis.Sci Rep. 2025 Jan 31;15(1):3925. doi: 10.1038/s41598-025-87438-1. Sci Rep. 2025. PMID: 39890881 Free PMC article.
-
Similarity-weighted entropy for quantifying genetic diversity in viral quasispecies.Virus Evol. 2025 Apr 26;11(1):veaf029. doi: 10.1093/ve/veaf029. eCollection 2025. Virus Evol. 2025. PMID: 40376429 Free PMC article.
-
Profiles of Circulating Exosomal miRNA in Patients with SAPHO Patients by High-Throughput Sequencing.Comb Chem High Throughput Screen. 2025;28(7):1193-1204. doi: 10.2174/0113862073289083240425114858. Comb Chem High Throughput Screen. 2025. PMID: 38752639
-
CPMI: comprehensive neighborhood-based perturbed mutual information for identifying critical states of complex biological processes.BMC Bioinformatics. 2024 Jun 15;25(1):215. doi: 10.1186/s12859-024-05836-0. BMC Bioinformatics. 2024. PMID: 38879513 Free PMC article.
References
MeSH terms
Substances
Grants and funding
- Nos. 61673008/National Natural Science Foundation of China
- No. 2019GGJS079/the Young Backbone Teacher Funding Scheme of Henan
- No. 212102310988/Key R & D and Promotion Special Program of Henan Province
- Grant Nos. 222102210053/the Key Science and Technology Research Project of Henan Province of China
- Grant Nos. 21A510003/the Key Scientific Research Project in Colleges and Universities of Henan Province of China
LinkOut - more resources
Full Text Sources

