SARS-CoV-2 shedding and evolution in patients who were immunocompromised during the omicron period: a multicentre, prospective analysis
- PMID: 38286131
- PMCID: PMC11849777
- DOI: 10.1016/S2666-5247(23)00336-1
SARS-CoV-2 shedding and evolution in patients who were immunocompromised during the omicron period: a multicentre, prospective analysis
Abstract
Background: Prolonged SARS-CoV-2 infections in people who are immunocompromised might predict or source the emergence of highly mutated variants. The types of immunosuppression placing patients at highest risk for prolonged infection have not been systematically investigated. We aimed to assess risk factors for prolonged SARS-CoV-2 infection and associated intrahost evolution.
Methods: In this multicentre, prospective analysis, participants were enrolled at five US medical centres. Eligible patients were aged 18 years or older, were SARS-CoV-2-positive in the previous 14 days, and had a moderately or severely immunocompromising condition or treatment. Nasal specimens were tested by real-time RT-PCR every 2-4 weeks until negative in consecutive specimens. Positive specimens underwent viral culture and whole genome sequencing. A Cox proportional hazards model was used to assess factors associated with duration of infection.
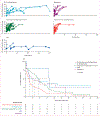
Findings: From April 11, 2022, to Oct 1, 2022, 156 patients began the enrolment process, of whom 150 were enrolled and included in the analyses. Participants had B-cell malignancy or anti-B-cell therapy (n=18), solid organ transplantation or haematopoietic stem-cell transplantation (HSCT; n=59), AIDS (n=5), non-B-cell malignancy (n=23), and autoimmune or autoinflammatory conditions (n=45). 38 (25%) participants were real-time RT-PCR-positive and 12 (8%) were culture-positive 21 days or longer after initial SARS-CoV-2 detection or illness onset. Compared with the group with autoimmune or autoinflammatory conditions, patients with B-cell dysfunction (adjusted hazard ratio 0·32 [95% CI 0·15-0·64]), solid organ transplantation or HSCT (0·60 [0·38-0·94]), and AIDS (0·28 [0·08-1·00]) had longer duration of infection, defined as time to last positive real-time RT-PCR test. There was no significant difference in the non-B-cell malignancy group (0·58 [0·31-1·09]). Consensus de novo spike mutations were identified in five individuals who were real-time RT-PCR-positive longer than 56 days; 14 (61%) of 23 were in the receptor-binding domain. Mutations shared by multiple individuals were rare (<5%) in global circulation.
Interpretation: In this cohort, prolonged replication-competent omicron SARS-CoV-2 infections were uncommon. Within-host evolutionary rates were similar across patients, but individuals with infections lasting longer than 56 days accumulated spike mutations, which were distinct from those seen globally. Populations at high risk should be targeted for repeated testing and treatment and monitored for the emergence of antiviral resistance.
Funding: US Centers for Disease Control and Prevention.
Copyright © 2023 The Author(s). Published by Elsevier Ltd.. All rights reserved.
Conflict of interest statement
Declaration of interests JDC reports receiving grants from the National Institutes of Health (NIH) and the Department of Defense, outside the submitted work. CGG reports grants from NIH, the Centers for Disease Control and Prevention (CDC), the Agency for Healthcare Research and Quality, the US Food and Drug Administration, and Campbell Alliance/Syneos Health, and consulting fees and participating on a data safety monitoring board for Merck, outside the submitted work. AEF reports a K08 award from NIH and participating on the Hennepin Health Research Institute Board of Directors, outside the submitted work. NH reports grants from Sanofi, Quidel, and Merck, outside the submitted work. ASL reports receiving grants from CDC, National Institute of Allergy and Infectious Diseases, Burroughs Wellcome Fund, and Flu Lab, and consulting fees from Roche, outside the submitted work. ETM reports receiving a grant from Merck, outside the submitted work. All other authors declare no competing interests.
Figures


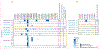
Update of
-
SARS-CoV-2 shedding and evolution in immunocompromised hosts during the Omicron period: a multicenter prospective analysis.medRxiv [Preprint]. 2023 Aug 24:2023.08.22.23294416. doi: 10.1101/2023.08.22.23294416. medRxiv. 2023. Update in: Lancet Microbe. 2024 Mar;5(3):e235-e246. doi: 10.1016/S2666-5247(23)00336-1. PMID: 37662226 Free PMC article. Updated. Preprint.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

