Genome-wide loss-of-function genetic screen identifies INSIG2 as the vulnerability of hepatitis B virus-integrated hepatoma cells
- PMID: 38287498
- PMCID: PMC10920982
- DOI: 10.1111/cas.16070
Genome-wide loss-of-function genetic screen identifies INSIG2 as the vulnerability of hepatitis B virus-integrated hepatoma cells
Abstract
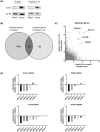
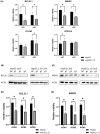
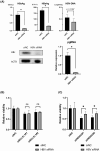
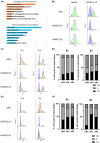
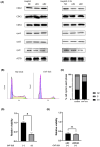
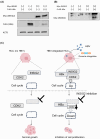
There are approximately 250 million people chronically infected with hepatitis B virus (HBV) worldwide. Although HBV is often integrated into the host genome and promotes hepatocarcinogenesis, vulnerability of HBV integration in liver cancer cells has not been clarified. The aim of our study is to identify vulnerability factors for HBV-associated hepatocarcinoma. Loss-of-function screening was undertaken in HepG2 and HBV-integrated HepG2.2.15 cells expressing SpCas9 using a pooled genome-wide clustered regularly interspaced short palindromic repeats (CRISPR) library. Genes whose guide RNA (gRNA) abundance significantly decreased in HepG2.2.15 cells but not in HepG2 cells were extracted using the MAGeCK algorithm. We identified four genes (BCL2L1, VPS37A, INSIG2, and CFLAR) that showed significant reductions of gRNA abundance and thus potentially involved in the vulnerability of HBV-integrated cancer cells. Among them, siRNA-mediated mRNA inhibition or CRISPR-mediated genetic deletion of INSIG2 significantly impaired cell proliferation in HepG2.2.15 cells but not in HepG2 cells. Its inhibitory effect was alleviated by cotransfection of siRNAs targeting HBV. INSIG2 inhibition suppressed the pathways related to cell cycle and DNA replication, downregulated cyclin-dependent kinase 2 (CDK2) levels, and delayed the G1 -to-S transition in HepG2.2.15 cells. CDK2 inhibitor suppressed cell cycle progression in HepG2.2.15 cells and INSIG2 inhibition did not suppress cell proliferation in the presence of CDK2 inhibitor. In conclusion, INSIG2 inhibition induced cell cycle arrest in HBV-integrated hepatoma cells in a CDK2-dependent manner, and thus INSIG2 might be a vulnerability factor for HBV-associated liver cancer.
Keywords: CDK2; CRISPR/Cas; HBV; liver cancer; pooled library screen.
© 2024 The Authors. Cancer Science published by John Wiley & Sons Australia, Ltd on behalf of Japanese Cancer Association.
Conflict of interest statement
The authors have no conflict of interest.
Figures
Similar articles
-
Targeted HBx gene editing by CRISPR/Cas9 system effectively reduces epithelial to mesenchymal transition and HBV replication in hepatoma cells.Liver Int. 2024 Feb;44(2):614-624. doi: 10.1111/liv.15805. Epub 2023 Dec 17. Liver Int. 2024. PMID: 38105495
-
Adenovirus Vectors Expressing Eight Multiplex Guide RNAs of CRISPR/Cas9 Efficiently Disrupted Diverse Hepatitis B Virus Gene Derived from Heterogeneous Patient.Int J Mol Sci. 2021 Sep 29;22(19):10570. doi: 10.3390/ijms221910570. Int J Mol Sci. 2021. PMID: 34638909 Free PMC article.
-
CRISPR/Cas9 System with Dual gRNAs Synergically Inhibit Hepatitis B Virus Replication.Discov Med. 2024 Jun;36(185):1169-1179. doi: 10.24976/Discov.Med.202436185.107. Discov Med. 2024. PMID: 38926103
-
Advances with using CRISPR/Cas-mediated gene editing to treat infections with hepatitis B virus and hepatitis C virus.Virus Res. 2018 Jan 15;244:311-320. doi: 10.1016/j.virusres.2017.01.003. Epub 2017 Jan 10. Virus Res. 2018. PMID: 28087399 Review.
-
The potential and challenges of CRISPR-Cas in eradication of hepatitis B virus covalently closed circular DNA.Virus Res. 2018 Jan 15;244:304-310. doi: 10.1016/j.virusres.2017.06.010. Epub 2017 Jun 13. Virus Res. 2018. PMID: 28627393 Review.
Cited by
-
Genome-wide CRISPR screen identifies BUB1 kinase as a druggable vulnerability in malignant pleural mesothelioma.Cell Death Dis. 2025 Apr 3;16(1):241. doi: 10.1038/s41419-025-07587-z. Cell Death Dis. 2025. PMID: 40180891 Free PMC article.
References
-
- Trepo C, Chan HLY, Lok A. Hepatitis B virus infection. Lancet. 2014;384:2053‐2063. - PubMed
-
- Organization WH . Guidelines for the prevention, care and treatment of persons with chronic hepatitis B infection. 2015. - PubMed
-
- Llovet JM, Kelley RK, Villanueva A, et al. Hepatocellular carcinoma. Nat Rev Dis Primers. 2021;7:6 - PubMed
MeSH terms
Substances
Grants and funding
- JP23ama221410/Japan Agency for Medical Research and Development
- JP23fk0210110/Japan Agency for Medical Research and Development
- JP23fk0210131/Japan Agency for Medical Research and Development
- JP23fk0310501/Japan Agency for Medical Research and Development
- JP23fk0310512/Japan Agency for Medical Research and Development
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

