MicroRNA-centered theranostics for pulmoprotection in critical COVID-19
- PMID: 38314095
- PMCID: PMC10834986
- DOI: 10.1016/j.omtn.2024.102118
MicroRNA-centered theranostics for pulmoprotection in critical COVID-19
Abstract
Elucidating the pathobiological mechanisms underlying post-acute pulmonary sequelae following SARS-CoV-2 infection is essential for early interventions and patient stratification. Here, we investigated the potential of microRNAs (miRNAs) as theranostic agents for pulmoprotection in critical illness survivors. Multicenter study including 172 ICU survivors. Diffusion impairment was defined as a lung-diffusing capacity for carbon monoxide (DLCO) <80% within 12 months postdischarge. A disease-associated 16-miRNA panel was quantified in plasma samples collected at ICU admission. Bioinformatic analyses were conducted using KEGG, Reactome, GTEx, and Drug-Gene Interaction databases. The results were validated using an external RNA-seq dataset. A 3-miRNA signature linked to diffusion impairment (miR-27a-3p, miR-93-5p, and miR-199a-5p) was identified using random forest. Levels of miR-93-5p and miR-199a-5p were independently associated with the outcome, improving patient classification provided by the electronic health record. The experimentally validated targets of these miRNAs exhibited enrichment across diverse pathways, with telomere length quantification in an additional set of samples (n = 83) supporting the role of cell senescence in sequelae. Analysis of an external dataset refined the pathobiological fingerprint of pulmonary sequelae. Gene-drug interaction analysis revealed four FDA-approved drugs. Overall, this study advances our understanding of lung recovery in postacute respiratory infections, highlighting the potential of miRNAs and their targets for pulmoprotection.
Keywords: MT: Novel therapeutic targets and biomarker development Special Issue; long COVID; microRNA; post-COVID syndrome; pulmoprotection; telomere; telomere length; theranostics.
© 2024 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
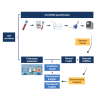

References
-
- Saunders C., Sperling S., Bendstrup E. A new paradigm is needed to explain long COVID. Lancet Respir. Med. 2023;11:e12–e13. - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

