A novel class of inhibitors that disrupts the stability of integrin heterodimers identified by CRISPR-tiling-instructed genetic screens
- PMID: 38316881
- PMCID: PMC10948361
- DOI: 10.1038/s41594-024-01211-y
A novel class of inhibitors that disrupts the stability of integrin heterodimers identified by CRISPR-tiling-instructed genetic screens
Abstract
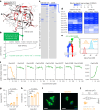
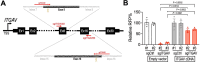
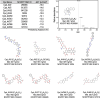
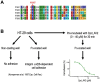
The plasma membrane is enriched for receptors and signaling proteins that are accessible from the extracellular space for pharmacological intervention. Here we conducted a series of CRISPR screens using human cell surface proteome and integrin family libraries in multiple cancer models. Our results identified ITGAV (integrin αV) and its heterodimer partner ITGB5 (integrin β5) as the essential integrin α/β pair for cancer cell expansion. High-density CRISPR gene tiling further pinpointed the integral pocket within the β-propeller domain of ITGAV for integrin αVβ5 dimerization. Combined with in silico compound docking, we developed a CRISPR-Tiling-Instructed Computer-Aided (CRISPR-TICA) pipeline for drug discovery and identified Cpd_AV2 as a lead inhibitor targeting the β-propeller central pocket of ITGAV. Cpd_AV2 treatment led to rapid uncoupling of integrin αVβ5 and cellular apoptosis, providing a unique class of therapeutic action that eliminates the integrin signaling via heterodimer dissociation. We also foresee the CRISPR-TICA approach to be an accessible method for future drug discovery studies.
© 2024. The Author(s).
Conflict of interest statement
J.C. is a scientific advisory board member of
Figures












References
MeSH terms
Grants and funding
- R01 CA236399/CA/NCI NIH HHS/United States
- R01 CA278050/CA/NCI NIH HHS/United States
- R00 CA197498/CA/NCI NIH HHS/United States
- K00 CA274649/CA/NCI NIH HHS/United States
- R01 CA255250/CA/NCI NIH HHS/United States
- R37 CA233691/CA/NCI NIH HHS/United States
- P30 CA033572/CA/NCI NIH HHS/United States
- R01 CA258778/CA/NCI NIH HHS/United States
- R01 CA243386/CA/NCI NIH HHS/United States
- F99 CA274649/CA/NCI NIH HHS/United States
- R01 CA236626/CA/NCI NIH HHS/United States
- U54 CA243124/CA/NCI NIH HHS/United States
- R01 CA271497/CA/NCI NIH HHS/United States
- R01 CA280389/CA/NCI NIH HHS/United States
- R01 GM117923/GM/NIGMS NIH HHS/United States
- R01 CA233922/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

