Tumor circadian clock strength influences metastatic potential and predicts patient prognosis in luminal A breast cancer
- PMID: 38319971
- PMCID: PMC10873596
- DOI: 10.1073/pnas.2311854121
Tumor circadian clock strength influences metastatic potential and predicts patient prognosis in luminal A breast cancer
Abstract
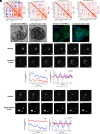
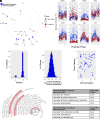
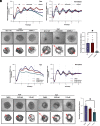
Studies in shift workers and model organisms link circadian disruption to breast cancer. However, molecular circadian rhythms in noncancerous and cancerous human breast tissues and their clinical relevance are largely unknown. We reconstructed rhythms informatically, integrating locally collected, time-stamped biopsies with public datasets. For noncancerous breast tissue, inflammatory, epithelial-mesenchymal transition (EMT), and estrogen responsiveness pathways show circadian modulation. Among tumors, clock correlation analysis demonstrates subtype-specific changes in circadian organization. Luminal A organoids and informatic ordering of luminal A samples exhibit continued, albeit dampened and reprogrammed rhythms. However, CYCLOPS magnitude, a measure of global rhythm strength, varied widely among luminal A samples. Cycling of EMT pathway genes was markedly increased in high-magnitude luminal A tumors. Surprisingly, patients with high-magnitude tumors had reduced 5-y survival. Correspondingly, 3D luminal A cultures show reduced invasion following molecular clock disruption. This study links subtype-specific circadian disruption in breast cancer to EMT, metastatic potential, and prognosis.
Keywords: breast cancer; circadian data ordering; circadian medicine; metastasis; prognosis.
Conflict of interest statement
Competing interests statement:J.B.H. is on the scientific advisory board for Synchronicity Pharma. The other authors declare no competing interest.
Figures


Comment in
-
Telling time in tumor samples reveals diversity of clock disruption.Proc Natl Acad Sci U S A. 2024 Mar 12;121(11):e2401496121. doi: 10.1073/pnas.2401496121. Epub 2024 Feb 29. Proc Natl Acad Sci U S A. 2024. PMID: 38422063 Free PMC article. No abstract available.
References
-
- Trayes K. P., Cokenakes S. E. H., Breast cancer treatment. Am. Fam. Phys. 104, 171–178 (2021). - PubMed
-
- Global Burden of Disease 2019 Cancer Collaboration. Cancer incidence, mortality, years of life lost, years lived with disability, and disability-adjusted life years for 29 cancer groups from 2010 to 2019: A systematic analysis for the global burden of disease study 2019. (2022). JAMA Oncol. 8, 420–444 (2022), 10.1001/jamaoncol.2021.6987. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

