Silica-coated magnetic particles for efficient RNA extraction for SARS-CoV-2 detection
- PMID: 38322940
- PMCID: PMC10844049
- DOI: 10.1016/j.heliyon.2024.e25377
Silica-coated magnetic particles for efficient RNA extraction for SARS-CoV-2 detection
Abstract
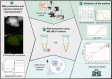
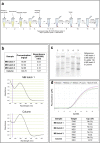
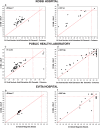
Molecular diagnostic methods to detect and quantify viral RNA in clinical samples rely on the purification of the genetic material prior to reverse transcription polymerase chain reaction (qRT-PCR). Due to the large number of samples processed in clinical laboratories, automation has become a necessity in order to increase method processivity and maximize throughput per unit of time. An attractive option for isolating viral RNA is based on the magnetic solid phase separation procedure (MSPS) using magnetic microparticles. This method offers the advantage over other alternative methods of making it possible to automate the process. In this study, we report the results of the MSPS method based on magnetic microparticles obtained by a simple synthesis process, to purify RNA from oro- and nasopharyngeal swab samples of patients suspected of COVID-19 provided by three diagnostic laboratories located in the Buenos Aires Province, Argentina. Magnetite nanoparticles of Fe3O4 (MNPs) were synthesized by the coprecipitation method and then coated with silica (SiO2) produced by hydrolysis of tetraethyl orthosilicate (TEOS). After preliminary tests on samples from the A549 human lung cell line and swabs, an extraction protocol was developed. The quantity and purity of the RNA obtained were determined by gel electrophoresis, spectrophotometry, and qRT-PCR. Tests on samples from naso- and oropharyngeal swabs were performed in order to validate the method for RNA purification in high-throughput SARS-CoV-2 diagnosis by qRT-PCR. The method was compared to the spin columns method and the automated method using commercial magnetic particles. The results show that the method developed is efficient for RNA extraction from nasal and oropharyngeal swab samples, and also comparable to other extraction methods in terms of sensitivity for SARS-CoV-2 detection. Of note, this procedure and reagents developed locally were intended to overcome the shortage of imported diagnostic supplies as the sudden spread of COVID-19 required unexpected quantities of nucleic acid isolation and diagnostic kits worldwide.
Keywords: Functionalized magnetic beads; MBs based protocol; Magnetic separation; Ribonucleic acid isolation; SARS-CoV-2 detection.
© 2024 The Authors. Published by Elsevier Ltd.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures



References
-
- Aislamiento social . Preventivo y obligatorio. 2020. https://www.boletinoficial.gob.ar/detalleAviso/primera/227042/20200320 Accessed: Jul. 11, 2023. [Online]. Available:
-
- de Salud Ministerio. Reporte diario vespertino nro 479. Situación de Covid-19 en Argentina. Ministerio de Salud, Argentina, Reporte diario vespertino. Dec. 2020:479. https://www.argentina.gob.ar/sites/default/files/30-12-20-reporte-vesper... [Online]. Available:
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

