Xanthomonas immunity proteins protect against the cis-toxic effects of their cognate T4SS effectors
- PMID: 38332152
- PMCID: PMC10933484
- DOI: 10.1038/s44319-024-00060-6
Xanthomonas immunity proteins protect against the cis-toxic effects of their cognate T4SS effectors
Abstract
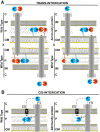
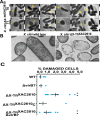
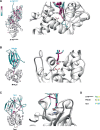
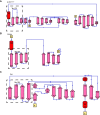
Many bacteria kill rival species by translocating toxic effectors into target cells. Effectors are often encoded along with cognate immunity proteins that could (i) protect against "friendly-fire" (trans-intoxication) from neighboring sister cells and/or (ii) protect against internal cis-intoxication (suicide). Here, we distinguish between these two mechanisms in the case of the bactericidal Xanthomonas citri Type IV Secretion System (X-T4SS). We use a set of X. citri mutants lacking multiple effector/immunity protein (X-Tfe/X-Tfi) pairs to show that X-Tfis are not absolutely required to protect against trans-intoxication by wild-type cells. Our investigation then focused on the in vivo function of the lysozyme-like effector X-TfeXAC2609 and its cognate immunity protein X-TfiXAC2610. In the absence of X-TfiXAC2610, we observe X-TfeXAC2609-dependent and X-T4SS-independent accumulation of damage in the X. citri cell envelope, cell death, and inhibition of biofilm formation. While immunity proteins in other systems have been shown to protect against attacks by sister cells (trans-intoxication), this is an example of an antibacterial secretion system in which the immunity proteins are dedicated to protecting cells against cis-intoxication.
Keywords: cis-intoxication; Bacterial Competition; Biofilm; Immunity Protein; Type IV Secretion System.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





Similar articles
-
Structural basis for effector recognition by an antibacterial type IV secretion system.Proc Natl Acad Sci U S A. 2022 Jan 4;119(1):e2112529119. doi: 10.1073/pnas.2112529119. Proc Natl Acad Sci U S A. 2022. PMID: 34983846 Free PMC article.
-
The opportunistic pathogen Stenotrophomonas maltophilia utilizes a type IV secretion system for interbacterial killing.PLoS Pathog. 2019 Sep 12;15(9):e1007651. doi: 10.1371/journal.ppat.1007651. eCollection 2019 Sep. PLoS Pathog. 2019. PMID: 31513674 Free PMC article.
-
Effectors of the Stenotrophomonas maltophilia Type IV Secretion System Mediate Killing of Clinical Isolates of Pseudomonas aeruginosa.mBio. 2021 Jun 29;12(3):e0150221. doi: 10.1128/mBio.01502-21. Epub 2021 Jun 29. mBio. 2021. PMID: 34182776 Free PMC article.
-
Bacteria-Killing Type IV Secretion Systems.Front Microbiol. 2019 May 21;10:1078. doi: 10.3389/fmicb.2019.01078. eCollection 2019. Front Microbiol. 2019. PMID: 31164878 Free PMC article. Review.
-
Type VI secretion system: a modular toolkit for bacterial dominance.Future Microbiol. 2019 Nov;14:1451-1463. doi: 10.2217/fmb-2019-0194. Epub 2019 Nov 13. Future Microbiol. 2019. PMID: 31718312 Review.
References
-
- Alegria MC, Souza DP, Andrade MO, Docena C, Khater L, Ramos CHI, da Silva ACR, Farah CS. Identification of new protein-protein interactions involving the products of the chromosome- and plasmid-encoded type IV secretion loci of the phytopathogen Xanthomonas axonopodis pv. citri. J Bacteriol. 2005;187:2315–2325. doi: 10.1128/JB.187.7.2315-2325.2005. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
- 2017/17303-7/Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP)
- 2018/09277-9/Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP)
- 2011/50521-1/Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP)
- 2014/04294-1/Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP)
- 2011/22571-4/Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP)
LinkOut - more resources
Full Text Sources

