Whole-Genome Sequencing Analyses Reveal the Whip-like Tail Formation, Innate Immune Evolution, and DNA Repair Mechanisms of Eupleurogrammus muticus
- PMID: 38338077
- PMCID: PMC10854985
- DOI: 10.3390/ani14030434
Whole-Genome Sequencing Analyses Reveal the Whip-like Tail Formation, Innate Immune Evolution, and DNA Repair Mechanisms of Eupleurogrammus muticus
Abstract
Smallhead hairtail (Eupleurogrammus muticus) is an important marine economic fish distributed along the northern Indian Ocean and the northwest Pacific coast; however, little is known about the mechanism of its genetic evolution. This study generated the first genome assembly of E. muticus at the chromosomal level using a combination of PacBio SMRT, Illumina Nova-Seq, and Hi-C technologies. The final assembled genome size was 709.27 Mb, with a contig N50 of 25.07 Mb, GC content of 40.81%, heterozygosity rate of 1.18%, and repetitive sequence rate of 35.43%. E. muticus genome contained 21,949 protein-coding genes (97.92% of the genes were functionally annotated) and 24 chromosomes. There were 143 expansion gene families, 708 contraction gene families, and 4888 positively selected genes in the genome. Based on the comparative genomic analyses, we screened several candidate genes and pathways related to whip-like tail formation, innate immunity, and DNA repair in E. muticus. These findings preliminarily reveal some molecular evolutionary mechanisms of E. muticus at the genomic level and provide important reference genomic data for the genetic studies of other trichiurids.
Keywords: Eupleurogrammus muticus; comparative genomics; genome sequencing; positive selection.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

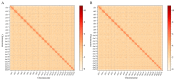
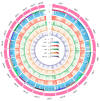
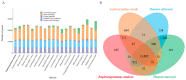



Similar articles
-
Chromosome-Level Genome Assembly of the Green Peafowl (Pavo muticus).Genome Biol Evol. 2022 Feb 4;14(2):evac015. doi: 10.1093/gbe/evac015. Genome Biol Evol. 2022. PMID: 35106558 Free PMC article.
-
Chromosome-Level Assembly of the Southern Rock Bream (Oplegnathus fasciatus) Genome Using PacBio and Hi-C Technologies.Front Genet. 2021 Dec 21;12:811798. doi: 10.3389/fgene.2021.811798. eCollection 2021. Front Genet. 2021. PMID: 34992639 Free PMC article. Review.
-
A Chromosomal-scale Reference Genome of the Kelp Grouper Epinephelus moara.Mar Biotechnol (NY). 2021 Feb;23(1):12-16. doi: 10.1007/s10126-020-10003-6. Epub 2020 Oct 8. Mar Biotechnol (NY). 2021. PMID: 33029658
-
Chromosome-level genome assembly of the East Asian common octopus (Octopus sinensis) using PacBio sequencing and Hi-C technology.Mol Ecol Resour. 2020 Nov;20(6):1572-1582. doi: 10.1111/1755-0998.13216. Epub 2020 Aug 20. Mol Ecol Resour. 2020. PMID: 32603549
-
Chromosome-level genome assembly of burbot (Lota lota) provides insights into the evolutionary adaptations in freshwater.Mol Ecol Resour. 2021 Aug;21(6):2022-2033. doi: 10.1111/1755-0998.13382. Epub 2021 Apr 3. Mol Ecol Resour. 2021. PMID: 33730415
References
-
- Froese R., Pauly D. FishBase. World Wide Web Electronic Publication. Version (10/2023) [(accessed on 3 November 2023)]. Available online: http://www.fishbase.org.
-
- Shao K.T. The Fish Database of Taiwan. [(accessed on 26 September 2023)]. Available online: http://fishdb.sinica.edu.tw.
-
- Nakamura I., Parin N.V. An annotated and illustrated catalogue of the Snake Mackerels, Snoeks, Escolars, Gemfishes, Sackfishes, Domine, Oilfish, Cutlassfishes, Scabbardfishes, Hairtails and Frostfishes known to date. FAO Fish. Synopis. 1993;125:61–107.
-
- Chen G.B., Li Y.Z., Zhao X.Y., Chen Y.Z., Jin X.S. Acoustic assessment of five groups commercial fish in South China Sea. Acta Oceanol. Sin. 2006;28:128–134.
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

