Analysis of Mutational Status of IGHV, and Cytokine Polymorphisms as Prognostic Factors in Chronic Lymphocytic Leukemia: The Romanian Experience
- PMID: 38339076
- PMCID: PMC10855205
- DOI: 10.3390/ijms25031799
Analysis of Mutational Status of IGHV, and Cytokine Polymorphisms as Prognostic Factors in Chronic Lymphocytic Leukemia: The Romanian Experience
Abstract
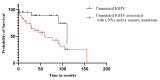
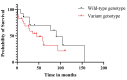
The aim of the current study was to assess the associations between genetic risk factors (such as the mutational status of the IGHV gene and polymorphisms of the IL-10 and TNF-α genes) and CLL risk, prognosis, and overall survival. Another goal of this study was to evaluate the multivariate effect of the combination of multiple genetic risk factors (mutational status of the IGHV gene, somatic mutations, DNA CNVs, and cytokine SNPs) on the clinical characteristics and survival of patients. A total of 125 CLL patients and 239 healthy controls were included for comparative SNP analysis. IL-10 (rs1800896 and rs1800872) and TNF-α (rs361525 and rs1800750) SNPs and haplotypes were not associated with CLL risk. The absence of hypermutation in the IGHV gene was shown to be of important prognostic value, being associated with short OS. Further individual risk factors for short OS were an age above 65 years at diagnosis and the presence of somatic mutations and/or CNVs. In our multivariable analysis, the presence of somatic mutations and the IL-10 rs1800872 variant allele, and the association of CNVs with the IL-10 rs1800896 variant allele, were identified as risk factors for short OS. Moreover, the OS in unmutated IGHV patients was additionally affected (decreased) by the presence of CNVs and/or somatic mutations. Similarly, IL-10 rs1800896 modulated the OS in unmutated IGHV patients with CNVs.
Keywords: CLL; IGHV; IL-10; TNF-α; cytokines; prognosis.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
Similar articles
-
Serum tumor necrosis factor-α and interleukin-10 levels as markers to predict outcome of patients with chronic lymphocytic leukemia in different risk groups defined by the IGHV mutation status.Arch Immunol Ther Exp (Warsz). 2012 Dec;60(6):477-86. doi: 10.1007/s00005-012-0197-7. Epub 2012 Sep 4. Arch Immunol Ther Exp (Warsz). 2012. PMID: 22945689
-
Cytokine rs361525, rs1800750, rs1800629, rs1800896, rs1800872, rs1800795, rs1800470, and rs2430561 SNPs in relation with prognostic factors in acute myeloid leukemia.Cancer Med. 2019 Sep;8(12):5492-5506. doi: 10.1002/cam4.2424. Epub 2019 Aug 1. Cancer Med. 2019. PMID: 31373163 Free PMC article.
-
The signs of negative selection in IGHV framework regions are associated with worse overall survival of chronic lymphocytic leukemia patients.Leuk Res. 2021 Nov;110:106686. doi: 10.1016/j.leukres.2021.106686. Epub 2021 Aug 25. Leuk Res. 2021. PMID: 34492598
-
[Impact of immunoglobulin gene somatic high mutation on prognosis of the chronic lymphocytic leukemia].Zhongguo Shi Yan Xue Ye Xue Za Zhi. 2009 Dec;17(6):1588-91. Zhongguo Shi Yan Xue Ye Xue Za Zhi. 2009. PMID: 20030953 Review. Chinese.
-
Searching for surrogates for IGHV mutations in chronic lymphocytic leukemia.Leuk Res. 2011 Nov;35(11):1432-5. doi: 10.1016/j.leukres.2011.07.020. Epub 2011 Aug 12. Leuk Res. 2011. PMID: 21840049 Review.
References
-
- Hiroki C.H., Amarante M.K., Petenuci D.L., Sakaguchi A.Y., Trigo F.C., Watanabe M.A.E., De Oliveira C.E.C. IL-10 Gene Polymorphism and Influence of Chemotherapy on Cytokine Plasma Levels in Childhood Acute Lymphoblastic Leukemia Patients. Blood Cells. Mol. Dis. 2015;55:168–172. doi: 10.1016/j.bcmd.2015.06.004. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

