RNF4 prevents genomic instability caused by chronic DNA under-replication
- PMID: 38340377
- PMCID: PMC10948022
- DOI: 10.1016/j.dnarep.2024.103646
RNF4 prevents genomic instability caused by chronic DNA under-replication
Abstract
Eukaryotic genome stability is maintained by a complex and diverse set of molecular processes. One class of enzymes that promotes proper DNA repair, replication and cell cycle progression comprises small ubiquitin-like modifier (SUMO)-targeted E3 ligases, or STUbLs. Previously, we reported a role for the budding yeast STUbL synthetically lethal with sgs1 (Slx) 5/8 in preventing G2/M-phase arrest in a minichromosome maintenance protein 10 (Mcm10)-deficient model of replication stress. Here, we extend these studies to human cells, examining the requirement for the human STUbL RING finger protein 4 (RNF4) in MCM10 mutant cancer cells. We find that MCM10 and RNF4 independently promote origin firing but regulate DNA synthesis epistatically and, unlike in yeast, the negative genetic interaction between RNF4 and MCM10 causes cells to accumulate in G1-phase. When MCM10 is deficient, RNF4 prevents excessive DNA under-replication at hard-to-replicate regions that results in large DNA copy number alterations and severely reduced viability. Overall, our findings highlight that STUbLs participate in species-specific mechanisms to maintain genome stability, and that human RNF4 is required for origin activation in the presence of chronic replication stress.
Keywords: Genome instability; MCM10; RNF4; Replication stress; Under-replicated DNA.
Copyright © 2024 The Authors. Published by Elsevier B.V. All rights reserved.
Conflict of interest statement
Declaration of Competing Interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures
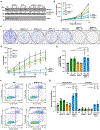


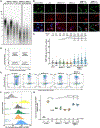

Similar articles
-
Acetyl transferase EP300 deficiency leads to chronic replication stress mediated by defective fork protection at stalled replication forks.bioRxiv [Preprint]. 2023 Apr 29:2023.04.29.538781. doi: 10.1101/2023.04.29.538781. bioRxiv. 2023. PMID: 37163075 Free PMC article. Preprint.
-
A critical threshold of MCM10 is required to maintain genome stability during differentiation of induced pluripotent stem cells into natural killer cells.Open Biol. 2024 Jan;14(1):230407. doi: 10.1098/rsob.230407. Epub 2024 Jan 24. Open Biol. 2024. PMID: 38262603 Free PMC article.
-
Disruption of SUMO-targeted ubiquitin ligases Slx5-Slx8/RNF4 alters RecQ-like helicase Sgs1/BLM localization in yeast and human cells.DNA Repair (Amst). 2015 Feb;26:1-14. doi: 10.1016/j.dnarep.2014.12.004. Epub 2014 Dec 26. DNA Repair (Amst). 2015. PMID: 25588990 Free PMC article.
-
SUMO-Targeted Ubiquitin Ligases and Their Functions in Maintaining Genome Stability.Int J Mol Sci. 2021 May 20;22(10):5391. doi: 10.3390/ijms22105391. Int J Mol Sci. 2021. PMID: 34065507 Free PMC article. Review.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2020 Jan 9;1(1):CD011535. doi: 10.1002/14651858.CD011535.pub3. Cochrane Database Syst Rev. 2020. Update in: Cochrane Database Syst Rev. 2021 Apr 19;4:CD011535. doi: 10.1002/14651858.CD011535.pub4. PMID: 31917873 Free PMC article. Updated.
Cited by
-
SUMO and the DNA damage response.Biochem Soc Trans. 2024 Apr 24;52(2):773-792. doi: 10.1042/BST20230862. Biochem Soc Trans. 2024. PMID: 38629643 Free PMC article. Review.
-
RNF135 promotes the stemness of breast cancer cells by ubiquitinating and degrading DDX58.Transl Oncol. 2025 Apr;54:102321. doi: 10.1016/j.tranon.2025.102321. Epub 2025 Feb 21. Transl Oncol. 2025. PMID: 39986189 Free PMC article.
-
Multifaceted roles of SUMO in DNA metabolism.Nucleus. 2024 Dec;15(1):2398450. doi: 10.1080/19491034.2024.2398450. Epub 2024 Sep 17. Nucleus. 2024. PMID: 39287196 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

