Inflammatory Fibroblast-Like Synoviocyte-Derived Exosomes Aggravate Osteoarthritis via Enhancing Macrophage Glycolysis
- PMID: 38342630
- PMCID: PMC11005727
- DOI: 10.1002/advs.202307338
Inflammatory Fibroblast-Like Synoviocyte-Derived Exosomes Aggravate Osteoarthritis via Enhancing Macrophage Glycolysis
Abstract
The severity of osteoarthritis (OA) and cartilage degeneration is highly associated with synovial inflammation. Although recent investigations have revealed a dysregulated crosstalk between fibroblast-like synoviocytes (FLSs) and macrophages in the pathogenesis of synovitis, limited knowledge is available regarding the involvement of exosomes. Here, increased exosome secretion is observed in FLSs from OA patients. Notably, internalization of inflammatory FLS-derived exosomes (inf-exo) can enhance the M1 polarization of macrophages, which further induces an OA-like phenotype in co-cultured chondrocytes. Intra-articular injection of inf-exo induces synovitis and exacerbates OA progression in murine models. In addition, it is demonstrated that inf-exo stimulation triggers the activation of glycolysis. Inhibition of glycolysis using 2-DG successfully attenuates excessive M1 polarization triggered by inf-exo. Mechanistically, HIF1A is identified as the determinant transcription factor, inhibition of which, both pharmacologically or genetically, relieves macrophage inflammation triggered by inf-exo-induced hyperglycolysis. Furthermore, in vivo administration of an HIF1A inhibitor alleviates experimental OA. The results provide novel insights into the involvement of FLS-derived exosomes in OA pathogenesis, suggesting that inf-exo-induced macrophage dysfunction represents an attractive target for OA therapy.
Keywords: HIF1A; exosomes; fibroblast‐like synoviocytes; glycolysis; macrophages; osteoarthritis; synovitis.
© 2024 The Authors. Advanced Science published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
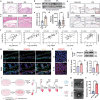

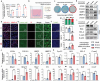



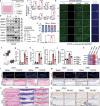
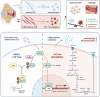
References
-
- Latourte A., Kloppenburg M., Richette P., Nat. Rev. Rheumatol. 2020, 16, 673. - PubMed
-
- Hügle T., Geurts J., Rheumatology 2017, 56, 1461. - PubMed
-
- Loeuille D., Chary‐Valckenaere I., Champigneulle J., Rat A. C., Toussaint F., Pinzano‐Watrin A., Goebel J. C., Mainard D., Blum A., Pourel J., Netter P., Gillet P., Arthritis Rheum. 2005, 52, 3492. - PubMed
MeSH terms
Grants and funding
- CXZX202214/Jiangsu Province Medical Innovation Center of Orthopedic Surgery
- 14380493/Fundamental Research Funds for the Central Universities
- 14380494/Fundamental Research Funds for the Central Universities
- 82272530/National Natural Science Foundation of China
- 82372459/National Natural Science Foundation of China
LinkOut - more resources
Full Text Sources
Medical
