Genomic vulnerability of a freshwater salmonid under climate change
- PMID: 38343776
- PMCID: PMC10853590
- DOI: 10.1111/eva.13602
Genomic vulnerability of a freshwater salmonid under climate change
Abstract
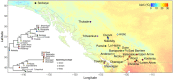
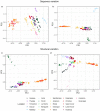
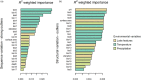
Understanding the adaptive potential of populations and species is pivotal for minimizing the loss of biodiversity in this era of rapid climate change. Adaptive potential has been estimated in various ways, including based on levels of standing genetic variation, presence of potentially beneficial alleles, and/or the severity of environmental change. Kokanee salmon, the non-migratory ecotype of sockeye salmon (Oncorhynchus nerka), is culturally and economically important and has already been impacted by the effects of climate change. To assess its climate vulnerability moving forward, we integrated analyses of standing genetic variation, genotype-environment associations, and climate modeling based on sequence and structural genomic variation from 224 whole genomes sampled from 22 lakes in British Columbia and Yukon (Canada). We found that variables for extreme temperatures, particularly warmer temperatures, had the most pervasive signature of selection in the genome and were the strongest predictors of levels of standing variation and of putatively adaptive genomic variation, both sequence and structural. Genomic offset estimates, a measure of climate vulnerability, were significantly correlated with higher increases in extreme warm temperatures, further highlighting the risk of summer heat waves that are predicted to increase in frequency in the future. Levels of standing genetic variation, an important metric for population viability and resilience, were not correlated with genomic offset. Nonetheless, our combined approach highlights the importance of integrating different sources of information and genomic data to formulate more comprehensive and accurate predictions on the vulnerability of populations and species to future climate change.
Keywords: Pacific salmon; climate vulnerability; local adaptation; standing variation; structural variants.
© 2023 The Authors. Evolutionary Applications published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no competing interest.
Figures


Similar articles
-
Population structure of sea-type and lake-type sockeye salmon and kokanee in the Fraser River and Columbia River drainages.PLoS One. 2017 Sep 8;12(9):e0183713. doi: 10.1371/journal.pone.0183713. eCollection 2017. PLoS One. 2017. PMID: 28886033 Free PMC article.
-
MOLECULAR GENETIC EVIDENCE FOR PARALLEL LIFE-HISTORY EVOLUTION WITHIN A PACIFIC SALMON (SOCKEYE SALMON AND KOKANEE, ONCORHYNCHUS NERKA).Evolution. 1996 Feb;50(1):401-416. doi: 10.1111/j.1558-5646.1996.tb04502.x. Evolution. 1996. PMID: 28568856
-
Snake River sockeye and Chinook salmon in a changing climate: Implications for upstream migration survival during recent extreme and future climates.PLoS One. 2020 Sep 30;15(9):e0238886. doi: 10.1371/journal.pone.0238886. eCollection 2020. PLoS One. 2020. PMID: 32997674 Free PMC article.
-
Dead fish swimming: a review of research on the early migration and high premature mortality in adult Fraser River sockeye salmon Oncorhynchus nerka.J Fish Biol. 2012 Jul;81(2):576-99. doi: 10.1111/j.1095-8649.2012.03360.x. Epub 2012 Jun 25. J Fish Biol. 2012. PMID: 22803725 Review.
-
Geospatial indicators of exposure, sensitivity, and adaptive capacity to assess neighbourhood variation in vulnerability to climate change-related health hazards.Environ Health. 2021 Mar 22;20(1):31. doi: 10.1186/s12940-021-00708-z. Environ Health. 2021. PMID: 33752667 Free PMC article.
Cited by
-
Assessment of future habitat suitability and ecological vulnerability of Collichthys at population and species level.BMC Ecol Evol. 2025 Jan 2;25(1):1. doi: 10.1186/s12862-024-02339-7. BMC Ecol Evol. 2025. PMID: 39743516 Free PMC article.
-
Advances in salmonid genetics-Insights from Coastwide and beyond.Evol Appl. 2024 Jun 17;17(6):e13732. doi: 10.1111/eva.13732. eCollection 2024 Jun. Evol Appl. 2024. PMID: 38887374 Free PMC article.
References
-
- Abdul‐Aziz, O. I. , Mantua, N. J. , & Myers, K. W. (2011). Potential climate change impacts on thermal habitats of Pacific salmon (Oncorhynchus spp.) in the North Pacific Ocean and adjacent seas. Canadian Journal of Fisheries and Aquatic Sciences, 68(9), 1660–1680.
-
- Akopyan, M. , Tigano, A. , Jacobs, A. , Wilder, A. P. , Baumann, H. , & Therkildsen, N. O. (2022). Comparative linkage mapping uncovers recombination suppression across massive chromosomal inversions associated with local adaptation in Atlantic silversides. Molecular Ecology, 31(12), 3323–3341. - PubMed
-
- Andrews, K. R. , Seaborn, T. , Egan, J. P. , Fagnan, M. W. , New, D. D. , Chen, Z. , Hohenlohe, P. A. , Waits, L. P. , Caudill, C. C. , & Narum, S. R. (2023). Whole genome resequencing identifies local adaptation associated with environmental variation for redband trout. Molecular Ecology, 32(4), 800–818. - PMC - PubMed
-
- Barrett, R. D. H. , & Schluter, D. (2008). Adaptation from standing genetic variation. Trends in Ecology & Evolution, 23(1), 38–44. - PubMed
LinkOut - more resources
Full Text Sources

