COVID-19 lateral flow test image classification using deep CNN and StyleGAN2
- PMID: 38348096
- PMCID: PMC10860423
- DOI: 10.3389/frai.2023.1235204
COVID-19 lateral flow test image classification using deep CNN and StyleGAN2
Abstract
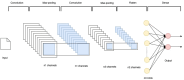
Introduction: Artificial intelligence (AI) in healthcare can enhance clinical workflows and diagnoses, particularly in large-scale operations like COVID-19 mass testing. This study presents a deep Convolutional Neural Network (CNN) model for automated COVID-19 RATD image classification.
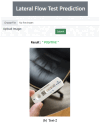
Methods: To address the absence of a RATD image dataset, we crowdsourced 900 real-world images focusing on positive and negative cases. Rigorous data augmentation and StyleGAN2-ADA generated simulated images to overcome dataset limitations and class imbalances.
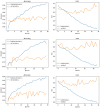
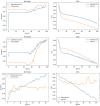
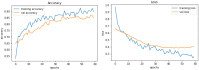
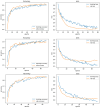
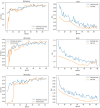
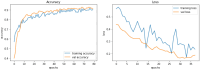
Results: The best CNN model achieved a 93% validation accuracy. Test accuracies were 88% for simulated datasets and 82% for real datasets. Augmenting simulated images during training did not significantly improve real-world test image performance but enhanced simulated test image performance.
Discussion: The findings of this study highlight the potential of the developed model in expediting COVID-19 testing processes and facilitating large-scale testing and tracking systems. The study also underscores the challenges in designing and developing such models, emphasizing the importance of addressing dataset limitations and class imbalances.
Conclusion: This research contributes to the deployment of large-scale testing and tracking systems, offering insights into the potential applications of AI in mitigating outbreaks similar to COVID-19. Future work could focus on refining the model and exploring its adaptability to other healthcare scenarios.
Keywords: SARS-CoV-2; StyleGAN2; convolutional neural network; deep learning; lateral flow test; transfer learning.
Copyright © 2024 Pannipulath Venugopal, Babu Saheer and Maktabdar Oghaz.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




Similar articles
-
Leveraging code-free deep learning for pill recognition in clinical settings: A multicenter, real-world study of performance across multiple platforms.Artif Intell Med. 2024 Apr;150:102844. doi: 10.1016/j.artmed.2024.102844. Epub 2024 Mar 13. Artif Intell Med. 2024. PMID: 38553153
-
An automated diagnosis and classification of COVID-19 from chest CT images using a transfer learning-based convolutional neural network.Comput Biol Med. 2022 May;144:105383. doi: 10.1016/j.compbiomed.2022.105383. Epub 2022 Mar 10. Comput Biol Med. 2022. PMID: 35290811 Free PMC article.
-
Automated image classification of chest X-rays of COVID-19 using deep transfer learning.Results Phys. 2021 Sep;28:104529. doi: 10.1016/j.rinp.2021.104529. Epub 2021 Jul 28. Results Phys. 2021. PMID: 34395185 Free PMC article.
-
Development and integration of VGG and dense transfer-learning systems supported with diverse lung images for discovery of the Coronavirus identity.Inform Med Unlocked. 2022;32:101004. doi: 10.1016/j.imu.2022.101004. Epub 2022 Jul 8. Inform Med Unlocked. 2022. PMID: 35822170 Free PMC article. Review.
-
Machine learning algorithms in microbial classification: a comparative analysis.Front Artif Intell. 2023 Oct 19;6:1200994. doi: 10.3389/frai.2023.1200994. eCollection 2023. Front Artif Intell. 2023. PMID: 37928448 Free PMC article. Review.
References
-
- Alazab M., Awajan A., Mesleh A., Abraham A., Jatana V., Alhyari S., et al. . (2020). COVID-19 prediction and detection using deep learning. Int. J. Comput. Inf. Syst. Ind. Manag. Appl. 12, 168–181.
-
- Appari N. V. L., Kanojia M. G. (2022). Soft computing and image processing techniques for COVID-19 prediction in lung CT scan images. Int. J. Hybrid Intell. Syst. 18, 111–131. 10.3233/HIS-220009 - DOI
-
- Arumugam S., Ma J., Macar U., Han G., McAulay K., Ingram D., et al. . (2021). Adaptable automated interpretation of rapid diagnostic tests using few-shot learning. medRxiv. [Preprint]. 10.1101/2021.06.23.21258927 - DOI
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

