Data-driven analysis and prediction of dynamic postprandial metabolic response to multiple dietary challenges using dynamic mode decomposition
- PMID: 38357465
- PMCID: PMC10865386
- DOI: 10.3389/fnut.2023.1304540
Data-driven analysis and prediction of dynamic postprandial metabolic response to multiple dietary challenges using dynamic mode decomposition
Abstract
Motivation: In the field of precision nutrition, predicting metabolic response to diet and identifying groups of differential responders are two highly desirable steps toward developing tailored dietary strategies. However, data analysis tools are currently lacking, especially for complex settings such as crossover studies with repeated measures.Current methods of analysis often rely on matrix or tensor decompositions, which are well suited for identifying differential responders but lacking in predictive power, or on dynamical systems modeling, which may be used for prediction but typically requires detailed mechanistic knowledge of the system under study. To remedy these shortcomings, we explored dynamic mode decomposition (DMD), which is a recent, data-driven method for deriving low-rank linear dynamical systems from high dimensional data.Combining the two recent developments "parametric DMD" (pDMD) and "DMD with control" (DMDc) enabled us to (i) integrate multiple dietary challenges, (ii) predict the dynamic response in all measured metabolites to new diets from only the metabolite baseline and dietary input, and (iii) identify inter-individual metabolic differences, i.e., metabotypes. To our knowledge, this is the first time DMD has been applied to analyze time-resolved metabolomics data.
Results: We demonstrate the potential of pDMDc in a crossover study setting. We could predict the metabolite response to unseen dietary exposures on both measured (R2 = 0.40) and simulated data of increasing size (= 0.65), as well as recover clusters of dynamic metabolite responses. We conclude that this method has potential for applications in personalized nutrition and could be useful in guiding metabolite response to target levels.
Availability and implementation: The measured data analyzed in this study can be provided upon reasonable request. The simulated data along with a MATLAB implementation of pDMDc is available at https://github.com/FraunhoferChalmersCentre/pDMDc.
Keywords: differential responders; dynamic mode decomposition; metabotypes; personalized nutrition; precision nutrition.
Copyright © 2024 Skantze, Jirstrand, Brunius, Sandberg, Landberg and Wallman.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures


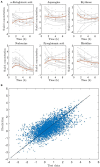





References
-
- Nicholson JK, Holmes E. Global systems biology and personalized healthcare solutions. Discov Med. (2006) 6:63–70. PMID: - PubMed
LinkOut - more resources
Full Text Sources

