Quantitative and qualitative mutational impact of ionizing radiation on normal cells
- PMID: 38359788
- PMCID: PMC10879144
- DOI: 10.1016/j.xgen.2024.100499
Quantitative and qualitative mutational impact of ionizing radiation on normal cells
Abstract
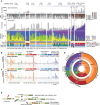
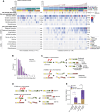
The comprehensive genomic impact of ionizing radiation (IR), a carcinogen, on healthy somatic cells remains unclear. Using large-scale whole-genome sequencing (WGS) of clones expanded from irradiated murine and human single cells, we revealed that IR induces a characteristic spectrum of short insertions or deletions (indels) and structural variations (SVs), including balanced inversions, translocations, composite SVs (deletion-insertion, deletion-inversion, and deletion-translocation composites), and complex genomic rearrangements (CGRs), including chromoplexy, chromothripsis, and SV by breakage-fusion-bridge cycles. Our findings suggest that 1 Gy IR exposure causes an average of 2.33 mutational events per Gb genome, comprising 2.15 indels, 0.17 SVs, and 0.01 CGRs, despite a high level of inter-cellular stochasticity. The mutational burden was dependent on total irradiation dose, regardless of dose rate or cell type. The findings were further validated in IR-induced secondary cancers and single cells without clonalization. Overall, our study highlights a comprehensive and clear picture of IR effects on normal mammalian genomes.
Keywords: carcinogen; clonalization; complex genomic rearrangement; ionizing radiation; irradiation dose; mutational signature; normal cell; single-base substitution; single-genome sequencing; structural variation.
Copyright © 2024 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests Y.S.J. is a genomic co-founder and chairman of Genome Insight.
Figures





References
MeSH terms
LinkOut - more resources
Full Text Sources

