Tissue-location-specific transcription programs drive tumor dependencies in colon cancer
- PMID: 38360902
- PMCID: PMC10869357
- DOI: 10.1038/s41467-024-45605-4
Tissue-location-specific transcription programs drive tumor dependencies in colon cancer
Abstract
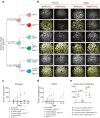
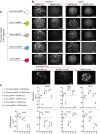
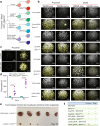
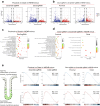
Cancers of the same tissue-type but in anatomically distinct locations exhibit different molecular dependencies for tumorigenesis. Proximal and distal colon cancers exemplify such characteristics, with BRAFV600E predominantly occurring in proximal colon cancers along with increased DNA methylation phenotype. Using mouse colon organoids, here we show that proximal and distal colon stem cells have distinct transcriptional programs that regulate stemness and differentiation. We identify that the homeobox transcription factor, CDX2, which is silenced by DNA methylation in proximal colon cancers, is a key mediator of the differential transcriptional programs. Cdx2-mediated proximal colon-specific transcriptional program concurrently is tumor suppressive, and Cdx2 loss sufficiently creates permissive state for BRAFV600E-driven transformation. Human proximal colon cancers with CDX2 downregulation showed similar transcriptional program as in mouse proximal organoids with Cdx2 loss. Developmental transcription factors, such as CDX2, are thus critical in maintaining tissue-location specific transcriptional programs that create tissue-type origin specific dependencies for tumor development.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

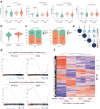
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

