Multiple steps of prion strain adaptation to a new host
- PMID: 38362022
- PMCID: PMC10867973
- DOI: 10.3389/fnins.2024.1329010
Multiple steps of prion strain adaptation to a new host
Abstract
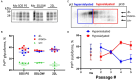
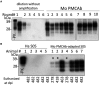
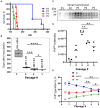
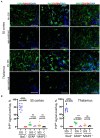
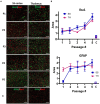
The transmission of prions across species is a critical aspect of their dissemination among mammalian hosts, including humans. This process often necessitates strain adaptation. In this study, we sought to investigate the mechanisms underlying prion adaptation while mitigating biases associated with the history of cross-species transmission of natural prion strains. To achieve this, we utilized the synthetic hamster prion strain S05. Propagation of S05 using mouse PrPC in Protein Misfolding Cyclic Amplification did not immediately overcome the species barrier. This finding underscores the involvement of factors beyond disparities in primary protein structures. Subsequently, we performed five serial passages to stabilize the incubation time to disease in mice. The levels of PrPSc increased with each passage, reaching a maximum at the third passage, and declining thereafter. This suggests that only the initial stage of adaptation is primarily driven by an acceleration in PrPSc replication. During the protracted adaptation to a new host, we observed significant alterations in the glycoform ratio and sialylation status of PrPSc N-glycans. These changes support the notion that qualitative modifications in PrPSc contribute to a more rapid disease progression. Furthermore, consistent with the decline in sialylation, a cue for "eat me" signaling, the newly adapted strain exhibited preferential colocalization with microglia. In contrast to PrPSc dynamics, the intensity of microglia activation continued to increase after the third passage in the new host. In summary, our study elucidates that the adaptation of a prion strain to a new host is a multi-step process driven by several factors.
Keywords: N-linked glycans; cross-species barrier; neurodegenerative diseases; prion; prion adaptation; prion diseases; prion strains; sialylation.
Copyright © 2024 Bocharova, Makarava, Pandit, Molesworth and Baskakov.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures
Update of
-
Multiple steps of prion strain adaptation to a new host.bioRxiv [Preprint]. 2023 Oct 27:2023.10.24.563743. doi: 10.1101/2023.10.24.563743. bioRxiv. 2023. Update in: Front Neurosci. 2024 Jan 31;18:1329010. doi: 10.3389/fnins.2024.1329010. PMID: 37961127 Free PMC article. Updated. Preprint.
References
-
- Asante E. A., Linehan J. M., Desbruslais M., Joiner S., Gowland I., Wood A. L., et al. (2002). BSE prions propagate as either variant CJD-like or sporadic CJD-like prion stains in transgenic mice expressing human prion protein. EMBO J. 21, 6358–6366. doi: 10.1093/emboj/cdf653, PMID: - DOI - PMC - PubMed
-
- Bartz J. C., Bessen R. A., McKenzie D., Marsh R. F., Aiken J. M. (2000). Adaptation and selection of prion protein strain conformations following interspecies transmission of transmissible mink encephalopathy. J. Virol. 74, 5542–5547. doi: 10.1128/JVI.74.12.5542-5547.2000, PMID: - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

