Whole-genome sequencing and comparative genomic analyses of the medicinal fungus Sanguinoderma infundibulare in Ganodermataceae
- PMID: 38366555
- PMCID: PMC10989896
- DOI: 10.1093/g3journal/jkae005
Whole-genome sequencing and comparative genomic analyses of the medicinal fungus Sanguinoderma infundibulare in Ganodermataceae
Abstract
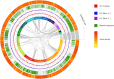
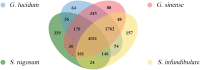
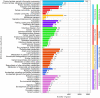
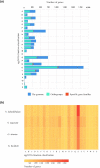
Sanguinoderma infundibulare is a newly discovered species of Ganodermataceae known to have high medicinal and ecological values. In this study, the whole-genome sequencing and comparative genomic analyses were conducted to further understand Ganodermataceae's genomic structural and functional characteristics. Using the Illumina NovaSeq and PacBio Sequel platforms, 88 scaffolds were assembled to obtain a 48.99-Mb high-quality genome of S. infundibulare. A total of 14,146 protein-coding genes were annotated in the whole genome, with 98.6% of complete benchmarking universal single-copy orthologs (BUSCO) scores. Comparative genomic analyses were conducted among S. infundibulare, Sanguinoderma rugosum, Ganoderma lucidum, and Ganoderma sinense to determine their intergeneric differences. The 4 species were found to share 4,011 orthogroups, and 24 specific gene families were detected in the genus Sanguinoderma. The gene families associated with carbohydrate esterase in S. infundibulare were significantly abundant, which was reported to be involved in hemicellulose degradation. One specific gene family in Sanguinoderma was annotated with siroheme synthase, which may be related to the typical characteristics of fresh pore surface changing to blood red when bruised. This study enriched the available genome data for the genus Sanguinoderma, elucidated the differences between Ganoderma and Sanguinoderma, and provided insights into the characteristics of the genome structure and function of S. infundibulare.
Keywords: Sanguinoderma; Ganodermataceae; comparative genomic analyses; medicinal fungi; next-generation sequencing.
© The Author(s) 2024. Published by Oxford University Press on behalf of The Genetics Society of America.
Conflict of interest statement
Conflicts of interest. The author(s) declare no conflicts of interest.
Figures




References
-
- Agrawal N, Verma P, Shahi SK. 2018. Degradation of polycyclic aromatic hydrocarbons (phenanthrene and pyrene) by the ligninolytic fungi Ganoderma lucidum isolated from the hardwood stump. Bioresour Bioprocess. 5(1):1–9. doi: 10.1186/s40643-018-0197-5. - DOI
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
