Phospholipids with two polyunsaturated fatty acyl tails promote ferroptosis
- PMID: 38366593
- PMCID: PMC10940216
- DOI: 10.1016/j.cell.2024.01.030
Phospholipids with two polyunsaturated fatty acyl tails promote ferroptosis
Abstract
Phospholipids containing a single polyunsaturated fatty acyl tail (PL-PUFA1s) are considered the driving force behind ferroptosis, whereas phospholipids with diacyl-PUFA tails (PL-PUFA2s) have been rarely characterized. Dietary lipids modulate ferroptosis, but the mechanisms governing lipid metabolism and ferroptosis sensitivity are not well understood. Our research revealed a significant accumulation of diacyl-PUFA phosphatidylcholines (PC-PUFA2s) following fatty acid or phospholipid treatments, correlating with cancer cell sensitivity to ferroptosis. Depletion of PC-PUFA2s occurred in aging and Huntington's disease brain tissue, linking it to ferroptosis. Notably, PC-PUFA2s interacted with the mitochondrial electron transport chain, generating reactive oxygen species (ROS) for initiating lipid peroxidation. Mitochondria-targeted antioxidants protected cells from PC-PUFA2-induced mitochondrial ROS (mtROS), lipid peroxidation, and cell death. These findings reveal a critical role for PC-PUFA2s in controlling mitochondria homeostasis and ferroptosis in various contexts and explain the ferroptosis-modulating mechanisms of free fatty acids. PC-PUFA2s may serve as diagnostic and therapeutic targets for modulating ferroptosis.
Keywords: PUFA; ROS; complex I; diacyl-PUFA phosphatidylcholine; electron transport chain; ferroptosis; lipids; mitochondria; phospholipid; polyunsaturated fatty acid.
Copyright © 2024 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests B.R.S. is an inventor on patents and patent applications involving ferroptosis; co-founded and serves as a consultant to ProJenX, Inc. and Exarta Therapeutics; holds equity in Sonata Therapeutics; serves as a consultant to Weatherwax Biotechnologies Corporation and Akin Gump Strauss Hauer & Feld LLP. C.T.B. is now the associate scientific director at Virgo Health.
Figures

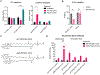




References
-
- Siegel MI, McConnell RT, Brent DA, Chandrasurin PISAL, Macfarlane RD, McNeal CJ (1982). The formation of diarachidonyl diglyceride by rat neutrophils. Mol. Pharmacol 21, 688–693. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

