Bacteriophage lambda site-specific recombination
- PMID: 38372210
- PMCID: PMC11096046
- DOI: 10.1111/mmi.15241
Bacteriophage lambda site-specific recombination
Abstract
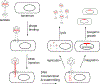
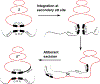
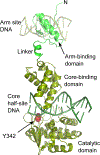
The site-specific recombination pathway of bacteriophage λ encompasses isoenergetic but highly directional and tightly regulated integrative and excisive reactions that integrate and excise the vial chromosome into and out of the bacterial chromosome. The reactions require 240 bp of phage DNA and 21 bp of bacterial DNA comprising 16 protein binding sites that are differentially used in each pathway by the phage-encoded Int and Xis proteins and the host-encoded integration host factor and factor for inversion stimulation proteins. Structures of higher-order protein-DNA complexes of the four-way Holliday junction recombination intermediates provided clarifying insights into the mechanisms, directionality, and regulation of these two pathways, which are tightly linked to the physiology of the bacterial host cell. Here we review our current understanding of the mechanisms responsible for regulating and executing λ site-specific recombination, with an emphasis on key studies completed over the last decade.
Keywords: DNA bending; Holliday junction; bacteriophage lambda; integration; site‐specific recombination; tyrosine recombinase; viral excision; viral integration.
© 2024 The Authors. Molecular Microbiology published by John Wiley & Sons Ltd.
Figures
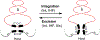

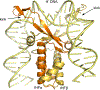

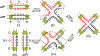

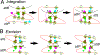
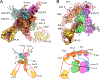
Similar articles
-
Mapping the λ Integrase bridges in the nucleoprotein Holliday junction intermediates of viral integrative and excisive recombination.Proc Natl Acad Sci U S A. 2014 Aug 26;111(34):12366-71. doi: 10.1073/pnas.1413007111. Epub 2014 Aug 11. Proc Natl Acad Sci U S A. 2014. PMID: 25114247 Free PMC article.
-
Nucleoprotein architectures regulating the directionality of viral integration and excision.Proc Natl Acad Sci U S A. 2014 Aug 26;111(34):12372-7. doi: 10.1073/pnas.1413019111. Epub 2014 Aug 11. Proc Natl Acad Sci U S A. 2014. PMID: 25114241 Free PMC article.
-
Structure of a Holliday junction complex reveals mechanisms governing a highly regulated DNA transaction.Elife. 2016 May 25;5:e14313. doi: 10.7554/eLife.14313. Elife. 2016. PMID: 27223329 Free PMC article.
-
DNA arms do the legwork to ensure the directionality of lambda site-specific recombination.Curr Opin Struct Biol. 2006 Feb;16(1):42-50. doi: 10.1016/j.sbi.2005.12.003. Epub 2005 Dec 20. Curr Opin Struct Biol. 2006. PMID: 16368232 Free PMC article. Review.
-
The λ Integrase Site-specific Recombination Pathway.Microbiol Spectr. 2015 Apr;3(2):MDNA3-0051-2014. doi: 10.1128/microbiolspec.MDNA3-0051-2014. Microbiol Spectr. 2015. PMID: 26104711 Free PMC article. Review.
Cited by
-
Large serine integrases utilise scavenged phage proteins as directionality cofactors.Nucleic Acids Res. 2025 Jan 24;53(3):gkaf050. doi: 10.1093/nar/gkaf050. Nucleic Acids Res. 2025. PMID: 39907112 Free PMC article.
-
Structural basis of directionality control in large serine integrases.bioRxiv [Preprint]. 2025 Jan 13:2025.01.03.631226. doi: 10.1101/2025.01.03.631226. bioRxiv. 2025. PMID: 39803483 Free PMC article. Preprint.
-
Dynamics in Cre-loxP site-specific recombination.Curr Opin Struct Biol. 2024 Oct;88:102878. doi: 10.1016/j.sbi.2024.102878. Epub 2024 Jul 18. Curr Opin Struct Biol. 2024. PMID: 39029281 Free PMC article. Review.
-
One-ended and two-ended breaks at nickase-broken replication forks.DNA Repair (Amst). 2024 Dec;144:103783. doi: 10.1016/j.dnarep.2024.103783. Epub 2024 Nov 4. DNA Repair (Amst). 2024. PMID: 39504607 Review.
References
-
- Abremski K. and Gottesman S. (1982) Purification of the bacteriophage lambda xis gene product required for lambda excisive recombination, The Journal of Biological Chemistry, 257, 9658–9662. - PubMed
-
- Aihara H. et al. (2003) A conformational switch controls the DNA cleavage activity of lambda integrase, Mol Cell, 12, 187–98. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

