An overview of actionable and potentially actionable TSC1 and TSC2 germline variants in an online Database
- PMID: 38373162
- PMCID: PMC10876083
- DOI: 10.1590/1678-4685-GMB-2023-0132
An overview of actionable and potentially actionable TSC1 and TSC2 germline variants in an online Database
Abstract
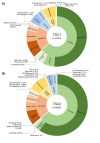
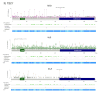
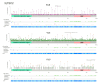
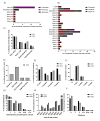
Tuberous Sclerosis Complex (TSC) is caused by loss of function germline variants in the TSC1 or TSC2 tumor suppressor genes. Genetic testing for the detection of pathogenic variants in either TSC1 or TSC2 was implemented as a diagnostic criterion for TSC. However, TSC molecular diagnosis can be challenging due to the absence of variant hotspots and the high number of variants described. This review aimed to perform an overview of TSC1/2 variants submitted in the ClinVar database. Variants of uncertain significance (VUS), missense and single nucleotide variants were the most frequent in clinical significance (37-40%), molecular consequence (37%-39%) and variation type (82%-83%) categories in ClinVar in TSC1 and TSC2 variants, respectively. Frameshift and nonsense VUS have potential for pathogenic reclassification if further functional and segregation studies were performed. Indeed, there were few functional assays deposited in the database and literature. In addition, we did not observe hotspots for variation and many variants presented conflicting submissions regarding clinical significance. This study underscored the importance of disseminating molecular diagnostic results in a public database to render the information largely accessible and promote accurate diagnosis. We encourage the performance of functional studies evaluating the pathogenicity of TSC1/2 variants.
Conflict of interest statement
Figures
Similar articles
-
Analysis of Genotypes and Phenotypes in Chinese Patients With Tuberous Sclerosis Complex Harboring Novel Variants of TSC1 and TSC2 Genes.Int J Genomics. 2025 May 8;2025:6963280. doi: 10.1155/ijog/6963280. eCollection 2025. Int J Genomics. 2025. PMID: 40376498 Free PMC article.
-
Comprehensive genetic and phenotype analysis of 95 individuals with mosaic tuberous sclerosis complex.Am J Hum Genet. 2023 Jun 1;110(6):979-988. doi: 10.1016/j.ajhg.2023.04.002. Epub 2023 May 3. Am J Hum Genet. 2023. PMID: 37141891 Free PMC article.
-
Comparison of the functional and structural characteristics of rare TSC2 variants with clinical and genetic findings.Hum Mutat. 2020 Apr;41(4):759-773. doi: 10.1002/humu.23963. Epub 2019 Dec 19. Hum Mutat. 2020. PMID: 31799751 Free PMC article.
-
Comprehensive mutation analysis of TSC1 and TSC2-and phenotypic correlations in 150 families with tuberous sclerosis.Am J Hum Genet. 1999 May;64(5):1305-15. doi: 10.1086/302381. Am J Hum Genet. 1999. PMID: 10205261 Free PMC article. Review.
-
Sclerotic Bone Lesions as a Clue in the Diagnosis of Three Generations of Tuberous Sclerosis Complex: Case Report and Review of Literature.Pediatr Neurol. 2023 Nov;148:14-16. doi: 10.1016/j.pediatrneurol.2023.07.022. Epub 2023 Aug 3. Pediatr Neurol. 2023. PMID: 37634327 Review.
Cited by
-
Neuroendocrine Tumors: Germline Genetics and Hereditary Syndromes.Curr Treat Options Oncol. 2025 Jan;26(1):55-71. doi: 10.1007/s11864-024-01288-z. Epub 2025 Jan 17. Curr Treat Options Oncol. 2025. PMID: 39821711 Review.
-
Tuberous Sclerosis Complex: A Case Series from a Romanian Genetics Center and a Review of the Literature.J Clin Med. 2025 Apr 25;14(9):2974. doi: 10.3390/jcm14092974. J Clin Med. 2025. PMID: 40364023 Free PMC article.
References
-
- Ali M, Girimaji SC, Markandaya M, Shukla AK, Sacchidanand S, Kumar A. Mutation and polymorphism analysis of TSC1 and TSC2 genes in Indian patients with Tuberous Sclerosis Complex. Acta Neurol Scand. 2005;111:54–63. - PubMed
-
- Amendola LM, Jarvik GP, Leo MC, Mclaughlin HM, Akkari Y, Amaral MD, Berg JS, Biswas S, Bowling KM, Conlin LK, et al. Performance of ACMG-AMP variant-interpretation guidelines among nine laboratories in the clinical sequencing exploratory research consortium. Am J Hum Genet. 2016;99:247. - PMC - PubMed
-
- Au KS, Williams AT, Roach ES, Batchelor L, Sparagana SP, Delgado MR, Wheless JW, Baumgartner JE, Roa BB, Wilson CM, et al. Genotype/phenotype correlation in 325 individuals referred for a diagnosis of Tuberous Sclerosis Complex in the United States. Genet Med. 2007;9:88–100. - PubMed
-
- BoronaT S, Caruso P, AuladelL M, Van Eeghen A, Thiele EA. Arachnoid cysts in Tuberous Sclerosis Complex. Brain Dev. 2014;36:801–806. - PubMed
Internet Resources
-
- cBioPortal for Cancer Genomics (cBioPortal) [04 January 2023]. cBioPortal for Cancer Genomics (cBioPortal), https://www.cbioportal.org/mutation_mapper .
-
- National Center for Biotechnology Information (NCBI) [09 December 2022]. National Center for Biotechnology Information (NCBI), https://www.ncbi.nlm.nih.gov/
-
- VarSome The Human Genomics Community (VarSome) [26 January 2023]. VarSome The Human Genomics Community (VarSome), https://varsome.com/variant/hg38/chr9%3A132902604%3AC%3AT .
LinkOut - more resources
Full Text Sources

