Loss of symbiotic and increase of virulent bacteria through microbial networks in Lynch syndrome colon carcinogenesis
- PMID: 38375206
- PMCID: PMC10876293
- DOI: 10.3389/fonc.2023.1313735
Loss of symbiotic and increase of virulent bacteria through microbial networks in Lynch syndrome colon carcinogenesis
Abstract
Purpose: Through a pilot study, we performed whole gut metagenomic analysis in 17 Lynch syndrome (LS) families, including colorectal cancer (CRC) patients and their healthy first-degree relatives. In a second asymptomatic LS cohort (n=150) undergoing colonoscopy-screening program, individuals with early precancerous lesions were compared to those with a normal colonoscopy. Since bacteria are organized into different networks within the microbiota, we compared related network structures in patients and controls.
Experimental design: Fecal prokaryote DNA was extracted prior to colonoscopy for whole metagenome (n=34, pilot study) or 16s rRNA sequencing (validation study). We characterized bacteria taxonomy using Diamond/MEGAN6 and DADA2 pipelines and performed differential abundances using Shaman website. We constructed networks using SparCC inference tools and validated the construction's accuracy by performing qPCR on selected bacteria.
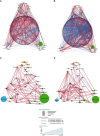
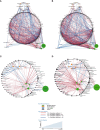
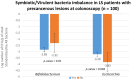
Results: Significant differences in bacterial communities in LS-CRC patients were identified, with an enrichment of virulent bacteria and a depletion of symbionts compared to their first-degree relatives. Bacteria taxa in LS asymptomatic individuals with colonic precancerous lesions (n=79) were significantly different compared to healthy individuals (n=71). The main bacterial network structures, constructed based on bacteria-bacteria correlations in CRC (pilot study) and in asymptomatic precancerous patients (validation-study), showed a different pattern than in controls. It was characterized by virulent/symbiotic co-exclusion in both studies and illustrated (validation study) by a higher Escherichia/Bifidobacterium ratio, as assessed by qPCR.
Conclusion: Enhanced fecal virulent/symbiotic bacteria ratios influence bacterial network structures. As an early event in colon carcinogenesis, these ratios can be used to identify asymptomatic LS individual with a higher risk of CRC.
Keywords: Lynch syndrome; cancer; colorectal cancer; gut microbiota; microbial network inference.
Copyright © 2024 Sadeghi, Mestivier, Carbonnelle, Benamouzig, Khazaie and Sobhani.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

