Reciprocal activity of AgRP and POMC neurons governs coordinated control of feeding and metabolism
- PMID: 38378998
- PMCID: PMC10963273
- DOI: 10.1038/s42255-024-00987-z
Reciprocal activity of AgRP and POMC neurons governs coordinated control of feeding and metabolism
Abstract
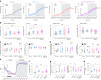
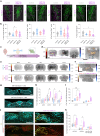
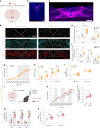
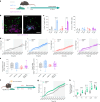
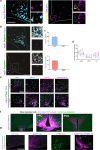
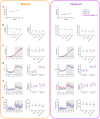
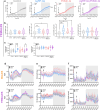
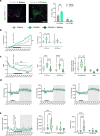
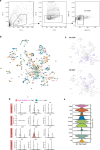
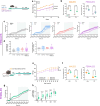
Agouti-related peptide (AgRP)-expressing and proopiomelanocortin (POMC)-expressing neurons reciprocally regulate food intake. Here, we combine non-interacting recombinases to simultaneously express functionally opposing chemogenetic receptors in AgRP and POMC neurons for comparing metabolic responses in male and female mice with simultaneous activation of AgRP and inhibition of POMC neurons with isolated activation of AgRP neurons or isolated inhibition of POMC neurons. We show that food intake is regulated by the additive effect of AgRP neuron activation and POMC neuron inhibition, while systemic insulin sensitivity and gluconeogenesis are differentially modulated by isolated-versus-simultaneous regulation of AgRP and POMC neurons. We identify a neurocircuit engaging Npy1R-expressing neurons in the paraventricular nucleus of the hypothalamus, where activated AgRP neurons and inhibited POMC neurons cooperate to promote food consumption and activate Th+ neurons in the nucleus tractus solitarii. Collectively, these results unveil how food intake is precisely regulated by the simultaneous bidirectional interplay between AgRP and POMC neurocircuits.
© 2024. The Author(s).
Conflict of interest statement
J.C.B. is cofounder and shareholder of Cerapeutix and has received research funding through collaborations with Sanofi Aventis and Novo Nordisk, and consultancy fees from Eli Lilly and Company and Novo Nordisk, which did not affect the content of this article. The other authors declare no competing interests.
Figures






References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

