Primary liver cancer classification from routine tumour biopsy using weakly supervised deep learning
- PMID: 38379584
- PMCID: PMC10877109
- DOI: 10.1016/j.jhepr.2024.101008
Primary liver cancer classification from routine tumour biopsy using weakly supervised deep learning
Abstract
Background & aims: The diagnosis of primary liver cancers (PLCs) can be challenging, especially on biopsies and for combined hepatocellular-cholangiocarcinoma (cHCC-CCA). We automatically classified PLCs on routine-stained biopsies using a weakly supervised learning method.
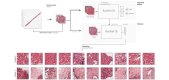
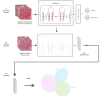
Method: We selected 166 PLC biopsies divided into training, internal and external validation sets: 90, 29 and 47 samples, respectively. Two liver pathologists reviewed each whole-slide hematein eosin saffron (HES)-stained image (WSI). After annotating the tumour/non-tumour areas, tiles of 256x256 pixels were extracted from the WSIs and used to train a ResNet18 neural network. The tumour/non-tumour annotations served as labels during training, and the network's last convolutional layer was used to extract new tumour tile features. Without knowledge of the precise labels of the malignancies, we then applied an unsupervised clustering algorithm.
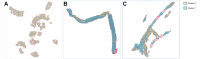
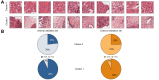
Results: Pathological review classified the training and validation sets into hepatocellular carcinoma (HCC, 33/90, 11/29 and 26/47), intrahepatic cholangiocarcinoma (iCCA, 28/90, 9/29 and 15/47), and cHCC-CCA (29/90, 9/29 and 6/47). In the two-cluster model, Clusters 0 and 1 contained mainly HCC and iCCA histological features. The diagnostic agreement between the pathological diagnosis and the two-cluster model predictions (major contingent) in the internal and external validation sets was 100% (11/11) and 96% (25/26) for HCC and 78% (7/9) and 87% (13/15) for iCCA, respectively. For cHCC-CCA, we observed a highly variable proportion of tiles from each cluster (cluster 0: 5-97%; cluster 1: 2-94%).
Conclusion: Our method applied to PLC HES biopsy could identify specific morphological features of HCC and iCCA. Although no specific features of cHCC-CCA were recognized, assessing the proportion of HCC and iCCA tiles within a slide could facilitate the identification of cHCC-CCA.
Impact and implications: The diagnosis of primary liver cancers can be challenging, especially on biopsies and for combined hepatocellular-cholangiocarcinoma (cHCC-CCA). We automatically classified primary liver cancers on routine-stained biopsies using a weakly supervised learning method. Our model identified specific features of hepatocellular carcinoma and intrahepatic cholangiocarcinoma. Despite no specific features of cHCC-CCA being recognized, the identification of hepatocellular carcinoma and intrahepatic cholangiocarcinoma tiles within a slide could facilitate the diagnosis of primary liver cancers, and particularly cHCC-CCA.
Keywords: Primary liver cancer; artificial intelligence; biopsy; histological slides; weakly supervised learning.
© 2024 The Author(s).
Conflict of interest statement
The authors declare no conflicts of interest that pertain to this work. Please refer to the accompanying ICMJE disclosure forms for further details.
Figures





References
-
- GLOBOCAN Cancer Fact Sheets Liver Cancers n.d. http://globocan.iarc.fr/old/FactSheets/cancers/liver-new.asp
LinkOut - more resources
Full Text Sources

