Exosomes define a local and systemic communication network in healthy pancreas and pancreatic ductal adenocarcinoma
- PMID: 38383468
- PMCID: PMC10881969
- DOI: 10.1038/s41467-024-45753-7
Exosomes define a local and systemic communication network in healthy pancreas and pancreatic ductal adenocarcinoma
Abstract
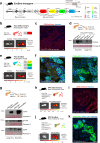
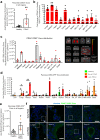
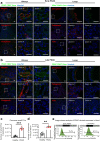
Pancreatic ductal adenocarcinoma (PDAC), a lethal disease, requires a grasp of its biology for effective therapies. Exosomes, implicated in cancer, are poorly understood in living systems. Here we use the genetically engineered mouse model (ExoBow) to map the spatiotemporal distribution of exosomes from healthy and PDAC pancreas in vivo to determine their biological significance. We show that, within the PDAC microenvironment, cancer cells establish preferential communication routes through exosomes with cancer associated fibroblasts and endothelial cells. The latter being a conserved event in the healthy pancreas. Inhibiting exosomes secretion in both scenarios enhances angiogenesis, underscoring their contribution to vascularization and to cancer. Inter-organ communication is significantly increased in PDAC with specific organs as most frequent targets of exosomes communication occurring in health with the thymus, bone-marrow, brain, and intestines, and in PDAC with the kidneys, lungs and thymus. In sum, we find that exosomes mediate an organized intra- and inter- pancreas communication network with modulatory effects in vivo.
© 2024. The Author(s).
Conflict of interest statement
S.A.M. holds patents in the area of EVs biology (miRNA biogenesis in exosomes for diagnosis and therapy, publication number 20200255831; use of exosomes for the treatment of disease, patent number 10959952; Analysis of genomic DNA, RNA and proteins in exosomes for diagnosis and theranosis, publication number 20200200755). The other authors declare no competing interests.
Figures





References
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

